Examination of methods to separate overlapping metabolites at 7T
- PMID: 39344348
- PMCID: PMC11604845
- DOI: 10.1002/mrm.30293
Examination of methods to separate overlapping metabolites at 7T
Abstract
Purpose: Neurochemicals of interest quantified by MRS are often composites of overlapping signals. At higher field strengths (i.e., 7T), there is better separation of these signals. As the availability of higher field strengths is increasing, it is important to re-evaluate the separability of overlapping metabolite signals.
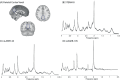
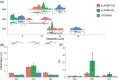
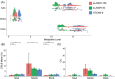
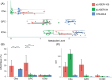
Methods: This study compares the ability of stimulated echo acquisition mode (STEAM-8; TE = 8 ms), short-TE semi-LASER (sLASER-34; TE = 34 ms), and long-TE semi-LASER (sLASER-105; TE = 105 ms) acquisitions to separate the commonly acquired neurochemicals at 7T (Glx, consisting of glutamate and glutamine; total N-acetyl aspartate, consisting of N-acetyl aspartate and N-acetylaspartylglutamate; total creatine, consisting of creatine and phosphocreatine; and total choline, consisting of choline, phosphocholine, and glycerophosphocholine).
Results: sLASER-34 produced the lowest fit errors for most neurochemicals; however, STEAM-8 had better within-subject reproducibility and required fewer subjects to detect a change between groups. However, this is dependent on the neurochemical of interest.
Conclusion: We recommend short-TE STEAM for separation of most standard neurochemicals at 7T over short-TE or long-TE sLASER.
Keywords: 7 T; STEAM; brain; high field; magnetic resonance spectroscopy; semi‐LASER.
© 2024 The Author(s). Magnetic Resonance in Medicine published by Wiley Periodicals LLC on behalf of International Society for Magnetic Resonance in Medicine.
Figures

References
MeSH terms
Substances
Grants and funding
- RGPIN-2017-03875/Natural Sciences and Engineering Research Council of Canada
- RGPIN-2020-05917/Natural Sciences and Engineering Research Council of Canada
- NFRFT 2022-00327/New Frontiers Research Fund of Canada
- Canadian Foundation for Innovation, John R. Evans Leaders Fund
- PJT-183715/CAPMC/ CIHR/Canada
LinkOut - more resources
Full Text Sources
Research Materials

