scEGG: an exogenous gene-guided clustering method for single-cell transcriptomic data
- PMID: 39344711
- PMCID: PMC11440090
- DOI: 10.1093/bib/bbae483
scEGG: an exogenous gene-guided clustering method for single-cell transcriptomic data
Abstract
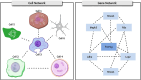
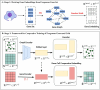
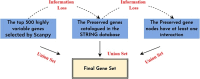
In recent years, there has been significant advancement in the field of single-cell data analysis, particularly in the development of clustering methods. Despite these advancements, most algorithms continue to focus primarily on analyzing the provided single-cell matrix data. However, within medical contexts, single-cell data often encompasses a wealth of exogenous information, such as gene networks. Overlooking this aspect could result in information loss and produce clustering outcomes lacking significant clinical relevance. To address this limitation, we introduce an innovative deep clustering method for single-cell data that leverages exogenous gene information to generate discriminative cell representations. Specifically, an attention-enhanced graph autoencoder has been developed to efficiently capture topological signal patterns among cells. Concurrently, a random walk on an exogenous protein-protein interaction network enabled the acquisition of the gene's embeddings. Ultimately, the clustering process entailed integrating and reconstructing gene-cell cooperative embeddings, which yielded a discriminative representation. Extensive experiments have demonstrated the effectiveness of the proposed method. This research provides enhanced insights into the characteristics of cells, thus laying the foundation for the early diagnosis and treatment of diseases. The datasets and code can be publicly accessed in the repository at https://github.com/DayuHuu/scEGG.
Keywords: Node2vec; clustering; deep learning; exogenous gene information; protein-protein interaction.
© The Author(s) 2024. Published by Oxford University Press.
Figures



Similar articles
-
scDFN: enhancing single-cell RNA-seq clustering with deep fusion networks.Brief Bioinform. 2024 Sep 23;25(6):bbae486. doi: 10.1093/bib/bbae486. Brief Bioinform. 2024. PMID: 39373051 Free PMC article.
-
scBGEDA: deep single-cell clustering analysis via a dual denoising autoencoder with bipartite graph ensemble clustering.Bioinformatics. 2023 Feb 14;39(2):btad075. doi: 10.1093/bioinformatics/btad075. Bioinformatics. 2023. PMID: 36734596 Free PMC article.
-
scMGATGRN: a multiview graph attention network-based method for inferring gene regulatory networks from single-cell transcriptomic data.Brief Bioinform. 2024 Sep 23;25(6):bbae526. doi: 10.1093/bib/bbae526. Brief Bioinform. 2024. PMID: 39417321 Free PMC article.
-
Machine learning and statistical methods for clustering single-cell RNA-sequencing data.Brief Bioinform. 2020 Jul 15;21(4):1209-1223. doi: 10.1093/bib/bbz063. Brief Bioinform. 2020. PMID: 31243426 Review.
-
Single-cell network biology for resolving cellular heterogeneity in human diseases.Exp Mol Med. 2020 Nov;52(11):1798-1808. doi: 10.1038/s12276-020-00528-0. Epub 2020 Nov 26. Exp Mol Med. 2020. PMID: 33244151 Free PMC article. Review.
Cited by
-
Graph neural networks for single-cell omics data: a review of approaches and applications.Brief Bioinform. 2025 Mar 4;26(2):bbaf109. doi: 10.1093/bib/bbaf109. Brief Bioinform. 2025. PMID: 40091193 Free PMC article.
References
-
- Peng L, Wang F, Wang Z. et al. .. Cell–cell communication inference and analysis in the tumour microenvironments from single-cell transcriptomics: data resources and computational strategies. Brief Bioinform 2022;23:bbac234. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

