Cpt1a Drives primed-to-naïve pluripotency transition through lipid remodeling
- PMID: 39349670
- PMCID: PMC11442460
- DOI: 10.1038/s42003-024-06874-3
Cpt1a Drives primed-to-naïve pluripotency transition through lipid remodeling
Abstract
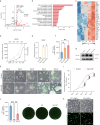
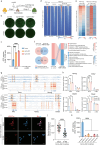
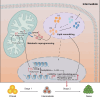
Metabolism has been implicated in cell fate determination, particularly through epigenetic modifications. Similarly, lipid remodeling also plays a role in regulating cell fate. Here, we present comprehensive lipidomics analysis during BMP4-driven primed to naive pluripotency transition or BiPNT and demonstrate that lipid remodeling plays an essential role. We further identify Cpt1a as a rate-limiting factor in BiPNT, driving lipid remodeling and metabolic reprogramming while simultaneously increasing intracellular acetyl-CoA levels and enhancing H3K27ac at chromatin open sites. Perturbation of BiPNT by histone acetylation inhibitors suppresses lipid remodeling and pluripotency transition. Together, our study suggests that lipid remodeling promotes pluripotency transitions and further regulates cell fate decisions, implicating Cpt1a as a critical regulator between primed-naive cell fate control.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

