Integrated GWAS, linkage, and transcriptome analysis to identify genetic loci and candidate genes for photoperiod sensitivity in maize
- PMID: 39351024
- PMCID: PMC11440433
- DOI: 10.3389/fpls.2024.1441288
Integrated GWAS, linkage, and transcriptome analysis to identify genetic loci and candidate genes for photoperiod sensitivity in maize
Abstract
Introduction: Maize photosensitivity and the control of flowering not only are important for reproduction, but also play pivotal roles in the processes of domestication and environmental adaptation, especially involving the utilization strategy of tropical maize in high-latitude regions.
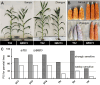
Methods: In this study, we used a linkage mapping population and an inbred association panel with the photoperiod sensitivity index (PSI) phenotyped under different environments and performed transcriptome analysis of T32 and QR273 between long-day and short-day conditions.
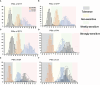
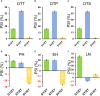
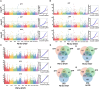
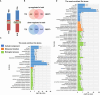
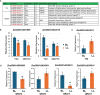
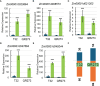
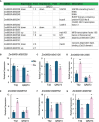
Results: The results showed that PSIs of days to tasseling (DTT), days to pollen shedding (DTP), and days to silking (DTS) indicated efficacious interactions with photoperiod sensitivity for maize latitude adaptation. A total of 48 quantitative trait loci (QTLs) and 252 quantitative trait nucleotides (QTNs) were detected using the linkage population and the inbred association panel. Thirteen candidate genes were identified by combining the genome-wide association study (GWAS) approach, linkage analysis, and transcriptome analysis, wherein five critical candidate genes, MYB163, bif1, burp8, CADR3, and Zm00001d050238, were significantly associated with photoperiod sensitivity.
Discussion: These results would provide much more abundant theoretical proofs to reveal the genetic basis of photoperiod sensitivity, which would be helpful to understand the genetic changes during domestication and improvement and contribute to reducing the barriers to use of tropical germplasm.
Keywords: GWAS; QTL; candidate gene; genetic loci; joint analysis; photoperiod sensitivity.
Copyright © 2024 Jiang, Guo, Wang, Tu, Liu, Guo, Wang, Zhu, Lu, Chen and Wu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
References
-
- Basu D., Wang W., Ma S., DeBrosse T., Poirier E., Emch K., et al. (2015). Two hydroxyproline galactosyltransferases, GALT5 and GALT2, function in arabinogalactan-protein glycosylation, growth and development in Arabidopsis. PloS One 10, e0125624. doi: 10.1371/journal.pone.0125624 - DOI - PMC - PubMed
-
- Birch C. J., Hammer G. L., Rickert K. G. (1998). Temperature and photoperiod sensitivity of development in five cultivars of maize (Zea mays L.) from emergence to tassel initiation. Field Crops Res. 55, 93–107. doi: 10.1016/S0378-4290(97)00062-2 - DOI
LinkOut - more resources
Full Text Sources

