Maintaining mitochondrial DNA copy number mitigates ROS-induced oocyte decline and female reproductive aging
- PMID: 39354016
- PMCID: PMC11445474
- DOI: 10.1038/s42003-024-06888-x
Maintaining mitochondrial DNA copy number mitigates ROS-induced oocyte decline and female reproductive aging
Abstract
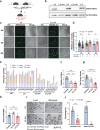
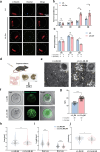
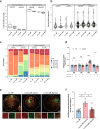
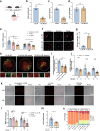
Oocytes play a crucial role in transmitting maternal mitochondrial DNA (mtDNA), essential for the continuation of species. However, the effects of mitochondrial reactive oxygen species (ROS) on mammalian oocyte maturation and mtDNA maintenance remain unclear. We investigated this by conditionally knocking out the Sod2 gene in primordial follicles, elevating mitochondrial matrix ROS levels from early oocyte stages. Our data indicates that reproductive aging in Sod2 conditional knockout females begins at 6 months, with oxidative stress impairing oocyte quality, particularly affecting OXPHOS complex II and mtDNA-encoded mRNA levels. Despite unchanged mtDNA mutation load, mtDNA copy numbers exhibited significant variations. Strikingly, reducing mtDNA copy numbers by reducing mtSSB protein, crucial for mtDNA replication, accelerated reproductive aging onset to three months, underscoring the critical role of mtDNA copy number maintenance under oxidative stress conditions. This research provides new insights into the relationship among mitochondrial ROS, mtDNA, and reproductive aging, offering potential strategies for delaying aging-related fertility decline.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Reproductive aging is associated with changes in oocyte mitochondrial dynamics, function, and mtDNA quantity.Maturitas. 2016 Nov;93:121-130. doi: 10.1016/j.maturitas.2016.06.015. Epub 2016 Jun 23. Maturitas. 2016. PMID: 27523387 Free PMC article.
-
The mitochondrial DNA copy number of cumulus granulosa cells may be related to the maturity of oocyte cytoplasm.Hum Reprod. 2020 May 1;35(5):1120-1129. doi: 10.1093/humrep/deaa085. Hum Reprod. 2020. PMID: 32358599
-
Maternal age and ovarian stimulation independently affect oocyte mtDNA copy number and cumulus cell gene expression in bovine clones.Hum Reprod. 2015 Jun;30(6):1410-20. doi: 10.1093/humrep/dev066. Epub 2015 Mar 27. Hum Reprod. 2015. PMID: 25820694
-
Mitochondrial unfolded protein response: a stress response with implications for fertility and reproductive aging.Fertil Steril. 2019 Feb;111(2):197-204. doi: 10.1016/j.fertnstert.2018.11.048. Fertil Steril. 2019. PMID: 30691623 Review.
-
Mitochondria: participation to infertility as source of energy and cause of senescence.Int J Biochem Cell Biol. 2014 Oct;55:60-4. doi: 10.1016/j.biocel.2014.08.011. Epub 2014 Aug 21. Int J Biochem Cell Biol. 2014. PMID: 25150832 Review.
Cited by
-
A Transcriptomics Approach to Unveil the Antioxidant Effects of Tryptophan on Oocyte Quality Under Oxidative Stress in Pigs.Biomolecules. 2025 Jun 30;15(7):949. doi: 10.3390/biom15070949. Biomolecules. 2025. PMID: 40723821 Free PMC article.
-
Mitochondrial dysfunction in oocytes: implications for fertility and ageing.J Ovarian Res. 2025 Aug 14;18(1):186. doi: 10.1186/s13048-025-01764-6. J Ovarian Res. 2025. PMID: 40814060 Free PMC article. Review.
-
Mitochondrial DNA Damage and Its Repair Mechanisms in Aging Oocytes.Int J Mol Sci. 2024 Dec 6;25(23):13144. doi: 10.3390/ijms252313144. Int J Mol Sci. 2024. PMID: 39684855 Free PMC article. Review.
-
The impact of mitochondrial dysfunction on ovarian aging.J Transl Med. 2025 Feb 20;23(1):211. doi: 10.1186/s12967-025-06223-w. J Transl Med. 2025. PMID: 39980008 Free PMC article. Review.
-
Endometrial Aging and Reproductive Decline: The Central Role of Mitochondrial Dysfunction.Int J Mol Sci. 2025 May 24;26(11):5060. doi: 10.3390/ijms26115060. Int J Mol Sci. 2025. PMID: 40507871 Free PMC article. Review.
References
-
- Wai, T., Teoli, D. & Shoubridge, E. A. The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat. Genet40, 1484–1488 (2008). - PubMed
-
- Cree, L. M. et al. A reduction of mitochondrial DNA molecules during embryogenesis explains the rapid segregation of genotypes. Nat. Genet40, 249–254 (2008). - PubMed
-
- Larsson, N.-G. et al. Mitochondrial transcription factor A is necessary for mtDNA maintance and embryogenesis in mice. Nat. Genet18, 231–236 (1998). - PubMed
-
- Broekmans, F. J., Soules, M. R. & Fauser, B. C. Ovarian Aging: Mechanisms and Clinical Consequences. Endocr. Rev.30, 465–493 (2009). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

