Quantitative evaluation of DNA damage repair dynamics to elucidate predictors of autism vs. cancer in individuals with germline PTEN variants
- PMID: 39356721
- PMCID: PMC11472915
- DOI: 10.1371/journal.pcbi.1012449
Quantitative evaluation of DNA damage repair dynamics to elucidate predictors of autism vs. cancer in individuals with germline PTEN variants
Abstract
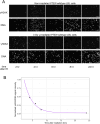
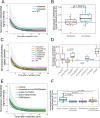
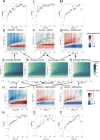
Persons with germline variants in the tumor suppressor gene phosphatase and tensin homolog, PTEN, are molecularly diagnosed with PTEN hamartoma tumor syndrome (PHTS). PHTS confers high risks of specific malignancies, and up to 23% of the patients are diagnosed with autism spectrum disorder (ASD) and/or developmental delay (DD). The accurate prediction of these two seemingly disparate phenotypes (cancer vs. ASD/DD) for PHTS at the individual level remains elusive despite the available statistical prevalence of specific phenotypes of the syndrome at the population level. The pleiotropy of the syndrome may, in part, be due to the alterations of the key multi-functions of PTEN. Maintenance of genome integrity is one of the key biological functions of PTEN, but no integrative studies have been conducted to quantify the DNA damage response (DDR) in individuals with PHTS and to relate to phenotypes and genotypes. In this study, we used 43 PHTS patient-derived lymphoblastoid cell lines (LCLs) to investigate the associations between DDR and PTEN genotypes and/or clinical phenotypes ASD/DD vs. cancer. The dynamics of DDR of γ-irradiated LCLs were analyzed using the exponential decay mathematical model to fit temporal changes in γH2AX levels which report the degree of DNA damage. We found that PTEN nonsense variants are associated with less efficient DNA damage repair ability resulting in higher DNA damage levels at 24 hours after irradiation compared to PTEN missense variants. Regarding PHTS phenotypes, LCLs from PHTS individuals with ASD/DD showed faster DNA damage repairing rate than those from patients without ASD/DD or cancer. We also applied the reaction-diffusion partial differential equation (PDE) mathematical model, a cell growth model with a DNA damage term, to accurately describe the DDR process in the LCLs. For each LCL, we can derive parameters of the PDE. Then we averaged the numerical results by PHTS phenotypes. By performing simple subtraction of two subgroup average results, we found that PHTS-ASD/DD is associated with higher live cell density at lower DNA damage level but lower cell density level at higher DNA damage level compared to LCLs from individuals with PHTS-cancer and PHTS-neither.
Copyright: © 2024 Wei et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
The mitochondrial genome as a modifier of autism versus cancer phenotypes in PTEN hamartoma tumor syndrome.HGG Adv. 2023 Apr 24;4(3):100199. doi: 10.1016/j.xhgg.2023.100199. eCollection 2023 Jul 13. HGG Adv. 2023. PMID: 37216009 Free PMC article.
-
Dynamics and structural stability effects of germline PTEN mutations associated with cancer versus autism phenotypes.J Biomol Struct Dyn. 2019 Apr;37(7):1766-1782. doi: 10.1080/07391102.2018.1465854. Epub 2018 May 14. J Biomol Struct Dyn. 2019. PMID: 29663862 Free PMC article.
-
Copy Number Variation and Clinical Outcomes in Patients With Germline PTEN Mutations.JAMA Netw Open. 2020 Jan 3;3(1):e1920415. doi: 10.1001/jamanetworkopen.2019.20415. JAMA Netw Open. 2020. PMID: 32003824 Free PMC article.
-
Cerebellar phenotypes in germline PTEN mutation carriers.Neuropathol Appl Neurobiol. 2024 Apr;50(2):e12970. doi: 10.1111/nan.12970. Neuropathol Appl Neurobiol. 2024. PMID: 38504418 Review.
-
PTEN hamartoma tumor syndrome.Handb Clin Neurol. 2015;132:129-37. doi: 10.1016/B978-0-444-62702-5.00009-3. Handb Clin Neurol. 2015. PMID: 26564076 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

