Genome-wide study of gene-by-sex interactions identifies risks for cleft palate
- PMID: 39361040
- PMCID: PMC12087800
- DOI: 10.1007/s00439-024-02704-y
Genome-wide study of gene-by-sex interactions identifies risks for cleft palate
Abstract
Structural birth defects affect 3-4% of all live births and, depending on the type, tend to manifest in a sex-biased manner. Orofacial clefts (OFCs) are the most common craniofacial structural birth defects and are often divided into cleft lip with or without cleft palate (CL/P) and cleft palate only (CP). Previous studies have found sex-specific risks for CL/P, but these risks have yet to be evaluated in CP. CL/P is more common in males and CP is more frequently observed in females, so we hypothesized there would also be sex-specific differences for CP. Using a trio-based cohort, we performed sex-stratified genome-wide association studies (GWAS) based on proband sex followed by a genome-wide gene-by-sex (G × S) interaction testing. There were 13 loci significant for G × S interactions, with the top finding in LTBP1 (RR = 3.37 [2.04-5.56], p = 1.93 × 10-6). LTBP1 plays a role in regulating TGF-β bioavailability, and knockdown in both mice and zebrafish lead to craniofacial anomalies. Further, there is evidence for differential expression of LTBP1 between males and females in both mice and humans. Therefore, we tested the association between the imputed genetically regulated gene expression of genes with significant G × S interactions and the CP phenotype. We found significant association for LTBP1 in cell cultured fibroblasts in female probands (p = 0.0013) but not in males. Taken altogether, we show there are sex-specific risks for CP that are otherwise undetectable in a combined sex cohort, and LTBP1 is a candidate risk gene, particularly in females.
© 2024. The Author(s), under exclusive licence to Springer-Verlag GmbH Germany, part of Springer Nature.
Conflict of interest statement
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Figures
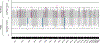
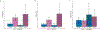

Update of
-
Genome-wide study of gene-by-sex interactions identifies risks for cleft palate.medRxiv [Preprint]. 2024 May 3:2024.05.01.24306701. doi: 10.1101/2024.05.01.24306701. medRxiv. 2024. Update in: Hum Genet. 2024 Nov;143(11):1341-1352. doi: 10.1007/s00439-024-02704-y. PMID: 38746184 Free PMC article. Updated. Preprint.
References
-
- Awotoye W, Comnick C, Pendleton C, Zeng E, Alade A, Mossey PA, Gowans LJJ, Eshete MA, Adeyemo WL, Naicker T, Adeleke C, Busch T, Li M, Petrin A, Olotu J, Hassan M, Pape J, Miller SE, Donkor P, Anand D, Lachke SA, Marazita ML, Adeyemo AA, Murray JC, Albokhari D, Sobreira N, and Butali A. 2022. ‘Genome-wide Gene-by-Sex Interaction Studies Identify Novel Nonsyndromic Orofacial Clefts Risk Locus’, Journal of Dental Research, 101: 465–72. - PMC - PubMed
-
- Barbeira Alvaro N., Dickinson Scott P., Bonazzola Rodrigo, Zheng Jiamao, Wheeler Heather E., Torres Jason M., Torstenson Eric S., Shah Kaanan P., Garcia Tzintzuni, Edwards Todd L., Stahl Eli A., Huckins Laura M., Aguet François, Ardlie Kristin G., Cummings Beryl B., Gelfand Ellen T., Getz Gad, Hadley Kane, Handsaker Robert E., Huang Katherine H., Kashin Seva, Karczewski Konrad J., Lek Monkol, Li Xiao, MacArthur Daniel G., Nedzel Jared L., Nguyen Duyen T., Noble Michael S., Segrè Ayellet V., Trowbridge Casandra A., Tukiainen Taru, Abell Nathan S., Balliu Brunilda, Barshir Ruth, Basha Omer, Battle Alexis, Bogu Gireesh K., Brown Andrew, Brown Christopher D., Castel Stephane E., Chen Lin S., Chiang Colby, Conrad Donald F., Damani Farhan N., Davis Joe R., Delaneau Olivier, Dermitzakis Emmanouil T., Engelhardt Barbara E., Eskin Eleazar, Ferreira Pedro G., Frésard Laure, Gamazon Eric R., Garrido-Martín Diego, Gewirtz Ariel D. H., Gliner Genna, Gloudemans Michael J., Guigo Roderic, Hall Ira M., Han Buhm, He Yuan, Hormozdiari Farhad, Howald Cedric, Jo Brian, Kang Eun Yong, Kim Yungil, Kim-Hellmuth Sarah, Lappalainen Tuuli, Li Gen, Li Xin, Liu Boxiang, Mangul Serghei, McCarthy Mark I., McDowell Ian C., Mohammadi Pejman, Monlong Jean, Montgomery Stephen B., Muñoz-Aguirre Manuel, Ndungu Anne W., Nobel Andrew B., Oliva Meritxell, Ongen Halit, Palowitch John J., Panousis Nikolaos, Papasaikas Panagiotis, Park YoSon, Parsana Princy, Payne Anthony J., Peterson Christine B., Quan Jie, Reverter Ferran, Sabatti Chiara, Saha Ashis, Sammeth Michael, Scott Alexandra J., Shabalin Andrey A., Sodaei Reza, Stephens Matthew, Stranger Barbara E., Strober Benjamin J., Sul Jae Hoon, Tsang Emily K., Urbut Sarah, van de Bunt Martijn, Wang Gao, Wen Xiaoquan, Wright Fred A., Xi Hualin S., Yeger-Lotem Esti, Zappala Zachary, Zaugg Judith B., Zhou Yi-Hui, Akey Joshua M., Bates Daniel, Chan Joanne, Chen Lin S., Claussnitzer Melina, Demanelis Kathryn, Diegel Morgan, Doherty Jennifer A., Feinberg Andrew P., Fernando Marian S., Halow Jessica, Hansen Kasper D., Haugen Eric, Hickey Peter F., Hou Lei, Jasmine Farzana, Jian Ruiqi, Jiang Lihua, Johnson Audra, Kaul Rajinder, Kellis Manolis, Kibriya Muhammad G., Lee Kristen, Jin Billy Li Qin Li, Li Xiao, Lin Jessica, Lin Shin, Linder Sandra, Linke Caroline, Liu Yaping, Maurano Matthew T., Molinie Benoit, Montgomery Stephen B., Nelson Jemma, Neri Fidencio J., Oliva Meritxell, Park Yongjin, Pierce Brandon L., Rinaldi Nicola J., Rizzardi Lindsay F., Sandstrom Richard, Skol Andrew, Smith Kevin S., Snyder Michael P., Stamatoyannopoulos John, Stranger Barbara E., Tang Hua, Tsang Emily K., Wang Li, Wang Meng, Nicholas Van Wittenberghe Fan Wu, Zhang Rui, Nierras Concepcion R., Branton Philip A., Carithers Latarsha J., Guan Ping, Moore Helen M., Rao Abhi, Vaught Jimmie B., Gould Sarah E., Lockart Nicole C., Martin Casey, Struewing Jeffery P., Volpi Simona, Addington Anjene M., Koester Susan E., Roger Little A, Brigham Lori E., Hasz Richard, Hunter Marcus, Johns Christopher, Johnson Mark, Kopen Gene, Leinweber William F., Lonsdale John T., Alisa McDonald Bernadette Mestichelli, Myer Kevin, Roe Brian, Salvatore Michael, Shad Saboor, Thomas Jeffrey A., Walters Gary, Washington Michael, Wheeler Joseph, Bridge Jason, Foster Barbara A., Gillard Bryan M., Karasik Ellen, Kumar Rachna, Miklos Mark, Moser Michael T., Jewell Scott D., Montroy Robert G., Rohrer Daniel C., Valley Dana R., Davis David A., Mash Deborah C., Undale Anita H., Smith Anna M., Tabor David E., Roche Nancy V., McLean Jeffrey A., Vatanian Negin, Robinson Karna L., Sobin Leslie, Barcus Mary E., Valentino Kimberly M., Qi Liqun, Hunter Steven, Hariharan Pushpa, Singh Shilpi, Um Ki Sung, Matose Takunda, Tomaszewski Maria M., Barker Laura K., Mosavel Maghboeba, Siminoff Laura A., Traino Heather M., Flicek Paul, Juettemann Thomas, Ruffier Magali, Sheppard Dan, Taylor Kieron, Trevanion Stephen J., Zerbino Daniel R., Craft Brian, Goldman Mary, Haeussler Maximilian, Kent W. James, Lee Christopher M., Paten Benedict, Rosenbloom Kate R., Vivian John, Zhu Jingchun, Nicolae Dan L., Cox Nancy J., Im Hae Kyung, G. TEx Consortium, Data Analysis Laboratory, Group Coordinating Center—Analysis Working, Group Statistical Methods groups—Analysis Working, GTEx groups Enhancing, N. I. H. Common Fund, Nih/Nci, Nih/NhgrI, Nih/Nimh, Nih/Nida, NDrI Biospecimen Collection Source Site—, P. C. I. Biospecimen Collection Source Site—r, VArI Biospecimen Core resource—, Bank Brain Bank repository—University of Miami Brain Endowment, Management Leidos Biomedical—Project, Elsi Study, Integration Genome Browser Data, Visualization—Ebi, Integration Genome Browser Data, and University of California Santa Cruz Visualization—Ucsc Genomics Institute. 2018. ‘Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics’, Nature Communications, 9: 1825. - PMC - PubMed
-
- Beaty TH, Ruczinski I, Murray JC, Marazita ML, Munger RG, Hetmanski JB, Murray T, Redett RJ, Fallin MD, Liang KY, Wu T, Patel PJ, Jin SC, Zhang TX, Schwender H, Wu-Chou YH, Chen PK, Chong SS, Cheah F, Yeow V, Ye X, Wang H, Huang S, Jabs EW, Shi B, Wilcox AJ, Lie RT, Jee SH, Christensen K, Doheny KF, Pugh EW, Ling H, and Scott AF. 2011. ‘Evidence for gene-environment interaction in a genome wide study of nonsyndromic cleft palate’, Genet Epidemiol, 35: 469–78. - PMC - PubMed
-
- Burdi AR, and Silvey RG. 1969. ‘Sexual differences in closure of the human palatal shelves’, Cleft Palate J, 6: 1–7. - PubMed
-
- Butali A, Mossey PA, Adeyemo WL, Eshete MA, Gowans LJJ, Busch TD, Jain D, Yu W, Huan L, Laurie CA, Laurie CC, Nelson S, Li M, Sanchez-Lara PA, Magee WP, Magee KS, Auslander A, Brindopke F, Kay DM, Caggana M, Romitti PA, Mills JL, Audu R, Onwuamah C, Oseni GO, Owais A, James O, Olaitan PB, Aregbesola BS, Braimah RO, Oginni FO, Oladele AO, Bello SA, Rhodes J, Shiang R, Donkor P, Obiri-Yeboah S, Arthur FKN, Twumasi P, Agbenorku P, Plange-Rhule G, Oti AA, Ogunlewe OM, Oladega AA, Adekunle AA, Erinoso AO, Adamson OO, Elufowoju AA, Ayelomi OI, Hailu T, Hailu A, Demissie Y, Derebew M, Eliason S, Romero-Bustillous M, Lo C, Park J, Desai S, Mohammed M, Abate F, Abdur-Rahman LO, Anand D, Saadi I, Oladugba AV, Lachke SA, Amendt BA, Rotimi CN, Marazita ML, Cornell RA, Murray JC, and Adeyemo AA. 2019. ‘Genomic analyses in African populations identify novel risk loci for cleft palate’, Hum Mol Genet, 28: 1038–51. - PMC - PubMed
MeSH terms
Substances
Grants and funding
- U54 GM133807/GM/NIGMS NIH HHS/United States
- R01 DE030342/DE/NIDCR NIH HHS/United States
- R01-DE028300/NH/NIH HHS/United States
- S21 MD001830/MD/NIMHD NIH HHS/United States
- X01-HG010835/NH/NIH HHS/United States
- R01 DE028300/DE/NIDCR NIH HHS/United States
- T32 GM008490/GM/NIGMS NIH HHS/United States
- R37-DE008559/NH/NIH HHS/United States
- U01 DD001226/DD/NCBDD CDC HHS/United States
- R00-DE024571/NH/NIH HHS/United States
- R01-DE016148/NH/NIH HHS/United States
- R37 DE008559/DE/NIDCR NIH HHS/United States
- R01 DE011931/DE/NIDCR NIH HHS/United States
- R01 DE014581/DE/NIDCR NIH HHS/United States
- R01-DE011931/NH/NIH HHS/United States
- R00 DE024571/DE/NIDCR NIH HHS/United States
- R01 DE016148/DE/NIDCR NIH HHS/United States
- T32-GM008490/NH/NIH HHS/United States
- F31 DE032588/DE/NIDCR NIH HHS/United States
- HHSN268201700006C/HL/NHLBI NIH HHS/United States
- K43 DE029427/DE/NIDCR NIH HHS/United States
- R01-DE014581/NH/NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

