GWAS and polygenic risk score of severe COVID-19 in Eastern Europe
- PMID: 39364016
- PMCID: PMC11446758
- DOI: 10.3389/fmed.2024.1409714
GWAS and polygenic risk score of severe COVID-19 in Eastern Europe
Abstract
Background: COVID-19 disease has infected more than 772 million people, leading to 7 million deaths. Although the severe course of COVID-19 can be prevented using appropriate treatments, effective interventions require a thorough research of the genetic factors involved in its pathogenesis.
Methods: We conducted a genome-wide association study (GWAS) on 7,124 individuals (comprising 6,400 controls who had mild to moderate COVID-19 and 724 cases with severe COVID-19). The inclusion criteria were acute respiratory distress syndrome (ARDS), acute respiratory failure (ARF) requiring respiratory support, or CT scans indicative of severe COVID-19 infection without any competing diseases. We also developed a polygenic risk score (PRS) model to identify individuals at high risk.
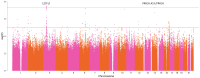
Results: We identified two genome-wide significant loci (P-value <5 × 10-8) and one locus with approximately genome-wide significance (P-value = 5.92 × 10-8-6.15 × 10-8). The most genome-wide significant variants were located in the leucine zipper transcription factor like 1 (LZTFL1) gene, which has been highlighted in several previous GWAS studies. Our PRS model results indicated that individuals in the top 10% group of the PRS had twice the risk of severe course of the disease compared to those at median risk [odds ratio = 2.18 (1.66, 2.86), P-value = 8.9 × 10-9].
Conclusion: We conducted one of the largest studies to date on the genetics of severe COVID-19 in an Eastern European cohort. Our results are consistent with previous research and will guide further epidemiologic studies on host genetics, as well as for the development of targeted treatments.
Keywords: COVID-19; GWAS; LZTFL1 gene; polygenic risk score; severe COVID-19.
Copyright © 2024 Kovalenko, Shaheen, Vergasova, Kamelin, Rubinova, Kharitonov, Kim, Plotnikov, Elmuratov, Borovkova, Storozheva, Solonin, Gilyazova, Mironov, Khusnutdinova, Petrikov, Ilinskaya, Ilinsky and Rakitko.
Conflict of interest statement
EKo, LS, EV, AKa, VR, DK, AKi, NP, and AR were employed by Genotek Ltd. AE was employed by Genetic Technologies Ltd. AI and VI were employed by Eligens SIA. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
-
- WHO coronavirus (COVID-19) dashboard. WHO Coronavirus (COVID-19) Dashboard with Vaccination Data. Available at: https://covid19.who.int/ (accessed November 30, 2023).
LinkOut - more resources
Full Text Sources

