Maintenance of hematopoietic stem cells by tyrosine-unphosphorylated STAT5 and JAK inhibition
- PMID: 39374575
- PMCID: PMC7617191
- DOI: 10.1182/bloodadvances.2024014046
Maintenance of hematopoietic stem cells by tyrosine-unphosphorylated STAT5 and JAK inhibition
Abstract
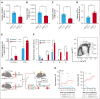
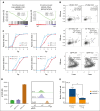
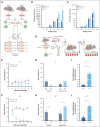
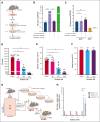
Adult hematopoietic stem cells (HSCs) are responsible for the lifelong production of blood and immune cells, a process regulated by extracellular cues, including cytokines. Many cytokines signal through the conserved Janus kinase (JAK)/signal transducer and activator of transcription (STAT) pathway in which tyrosine-phosphorylated STATs (pSTATs) function as transcription factors. STAT5 is a pivotal downstream mediator of several cytokines known to regulate hematopoiesis, but its function in the HSC compartment remains poorly understood. In this study, we show that STAT5-deficient HSCs exhibit an unusual phenotype, including reduced multilineage repopulation and self-renewal, combined with reduced exit from quiescence and increased differentiation. This was driven not only by the loss of canonical pSTAT5 signaling, but also by the loss of distinct transcriptional functions mediated by STAT5 that lack canonical tyrosine phosphorylation (uSTAT5). Consistent with this concept, expression of an unphosphorylatable STAT5 mutant constrained wild-type HSC differentiation, promoted their maintenance, and upregulated transcriptional programs associated with quiescence and stemness. The JAK1/2 inhibitor, ruxolitinib, which increased the uSTAT5:pSTAT5 ratio, had similar effects on murine HSC function; it constrained HSC differentiation and proliferation, promoted HSC maintenance, and upregulated transcriptional programs associated with stemness. Ruxolitinib also enhanced serial replating of normal human hematopoietic stem and progenitor cells (HSPCs), calreticulin-mutant murine HSCs, and HSPCs obtained from patients with myelofibrosis. Our results therefore reveal a previously unrecognized interplay between pSTAT5 and uSTAT5 in the control of HSC function and highlight JAK inhibition as a potential strategy for enhancing HSC function during ex vivo culture. Increased levels of uSTAT5 may also contribute to the failure of JAK inhibitors to eradicate myeloproliferative neoplasms.
© 2024 by The American Society of Hematology. This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/).
Conflict of interest statement
Conflict-of-interest disclosure: A.R.G. and J.L. report serving as consultants for Incyte. E.L. reports receiving research funds from GlaxoSmithKline and Commonwealth Serum Laboratories Behring. The remaining authors declare no competing financial interests.
Figures





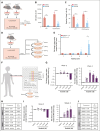
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

