The biogenesis, regulation and functions of transitive siRNA in plants
- PMID: 39376148
- PMCID: PMC11802348
- DOI: 10.3724/abbs.2024160
The biogenesis, regulation and functions of transitive siRNA in plants
Abstract
Small RNA (sRNA)-mediated RNA interference (RNAi) is a sequence-specific gene silencing mechanism that modulates gene expression in eukaryotes. As core molecules of RNAi, various sRNAs are encoded in the plant genome or derived from invading RNA molecules, and their biogenesis depends on distinct genetic pathways. Transitive small interfering RNAs (siRNAs), which are sRNAs produced from double-strand RNA (dsRNA) in a process that depends on RNA-dependent RNA polymerases (RDRs), can amplify and spread silencing signals to additional transcripts, thereby enabling a phenomenon termed "transitive RNAi". Members of this class of siRNAs function in various biological processes ranging from development to stress adaptation. In Arabidopsis thaliana, two RDRs participate in the generation of transitive siRNAs, acting cooperatively with various siRNA generation-related factors, such as the RNA-induced silencing complex (RISC) and aberrant RNAs. Transitive siRNAs are produced in diverse subcellular locations and structures under the control of various mechanisms, highlighting the intricacies of their biogenesis and functions. In this review, we discuss recent advances in understanding the molecular events of transitive siRNA biogenesis and its regulation, with a particular focus on factors involved in RDR recruitment. We aim to provide a comprehensive description of the generalized mechanism governing the biogenesis of transitive siRNAs. Additionally, we present an overview of the diverse biological functions of these siRNAs and raise some pressing questions in this area for further investigation.
Keywords: RDRs; siRNA; silencing; transitivity.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

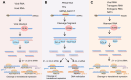
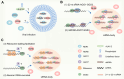
References
-
- Bologna NG, Voinnet O. The diversity, biogenesis, and activities of endogenous silencing small RNAs in Arabidopsis . Annu Rev Plant Biol. . 2014;65:473–503. doi: 10.1146/annurev-arplant-050213-035728. - DOI - PubMed
-
- Zhan J, Meyers BC. Plant small RNAs: their biogenesis, regulatory roles, and functions. Annu Rev Plant Biol. . 2023;74:21–51. doi: 10.1146/annurev-arplant-070122-035226. - DOI - PubMed
-
- Borges F, Martienssen RA. The expanding world of small RNAs in plants. Nat Rev Mol Cell Biol. . 2015;16:727–741. doi: 10.1038/nrm4085. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

