Frequency and spectrum of mutations in human sperm measured using duplex sequencing correlate with trio-based de novo mutation analyses
- PMID: 39379474
- PMCID: PMC11461794
- DOI: 10.1038/s41598-024-73587-2
Frequency and spectrum of mutations in human sperm measured using duplex sequencing correlate with trio-based de novo mutation analyses
Erratum in
-
Author Correction: Frequency and spectrum of mutations in human sperm measured using duplex sequencing correlate with trio-based de novo mutation analyses.Sci Rep. 2024 Dec 19;14(1):30571. doi: 10.1038/s41598-024-82618-x. Sci Rep. 2024. PMID: 39702384 Free PMC article. No abstract available.
Abstract
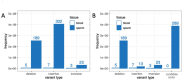
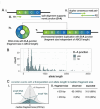
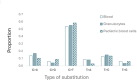
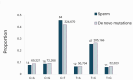
De novo mutations (DNMs) are drivers of genetic disorders. However, the study of DNMs is hampered by technological limitations preventing accurate quantification of ultra-rare mutations. Duplex Sequencing (DS) theoretically has < 1 error/billion base-pairs (bp). To determine the DS utility to quantify and characterize DNMs, we analyzed DNA from blood and spermatozoa from six healthy, 18-year-old Swedish men using the TwinStrand DS mutagenesis panel (48 kb spanning 20 genic and intergenic loci). The mean single nucleotide variant mutation frequency (MF) was 1.2 × 10- 7 per bp in blood and 2.5 × 10- 8 per bp in sperm, with the most common base substitution being C > T. Blood MF and substitution spectrum were similar to those reported in blood cells with an orthogonal method. The sperm MF was in the same order of magnitude and had a strikingly similar spectrum to DNMs from publicly available whole genome sequencing data from human pedigrees (1.2 × 10- 8 per bp). DS revealed much larger numbers of insertions and deletions in sperm over blood, driven by an abundance of putative extra-chromosomal circular DNAs. The study indicates the strong potential of DS to characterize human DNMs to inform factors that contribute to disease susceptibility and heritable genetic risks.
Keywords: De novo mutations; Duplex sequencing; Extrachromosomal circular DNA; Mutation frequency; Mutational spectrum; Sperm DNA mutations.
© 2024. The Author(s).
Conflict of interest statement
JA, CY, DLB, HS, MM, RP, AW and FM declare no competing interests. DF, JC, AG, JH, TS, DN, FYL and JS are employees and equity holders at TwinStrand Biosciences, Inc., or were during their contributions to this manuscript.
Figures


References
-
- Rosell, A. M. et al. Not the end of the odyssey: parental perceptions of whole exome sequencing (WES) in pediatric undiagnosed disorders. J. Genet. Couns.25, 1019–1031. 10.1007/s10897-016-9933-1 (2016). - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

