Human selenocysteine synthase, SEPSECS, has evolved to optimize binding of a tRNA-based substrate
- PMID: 39385655
- PMCID: PMC11602143
- DOI: 10.1093/nar/gkae875
Human selenocysteine synthase, SEPSECS, has evolved to optimize binding of a tRNA-based substrate
Abstract
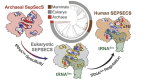
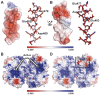
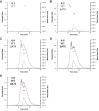
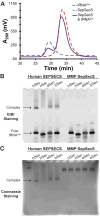
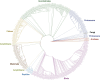
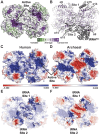
The evolution of the genetic code to incorporate selenocysteine (Sec) enabled the development of a selenoproteome in all domains of life. O-phosphoseryl-tRNASec selenium transferase (SepSecS) catalyzes the terminal reaction of Sec synthesis on tRNASec in archaea and eukaryotes. Despite harboring four equivalent active sites, human SEPSECS binds no more than two tRNASec molecules. Though, the basis for this asymmetry remains poorly understood. In humans, an acidic, C-terminal, α-helical extension precludes additional tRNA-binding events in two of the enzyme monomers, stabilizing the SEPSECS•tRNASec complex. However, the existence of a helix exclusively in vertebrates raised questions about the evolution of the tRNA-binding mechanism in SEPSECS and the origin of its C-terminal extension. Herein, using a comparative structural and phylogenetic analysis, we show that the tRNA-binding motifs in SEPSECS are poorly conserved across species. Consequently, in contrast to mammalian SEPSECS, the archaeal ortholog cannot bind unacylated tRNASec and requires an aminoacyl group. Moreover, the C-terminal α-helix 16 is a mammalian innovation, and its absence causes aggregation of the SEPSECS•tRNASec complex at low tRNA concentrations. Altogether, we propose SEPSECS evolved a tRNASec binding mechanism as a crucial functional and structural feature, allowing for additional levels of regulation of Sec and selenoprotein synthesis.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
Structural basis for the tRNA-dependent activation of the terminal complex of selenocysteine synthesis in humans.Nucleic Acids Res. 2023 May 8;51(8):4012-4026. doi: 10.1093/nar/gkad182. Nucleic Acids Res. 2023. PMID: 36929010 Free PMC article.
-
Structural asymmetry of the terminal catalytic complex in selenocysteine synthesis.J Biol Chem. 2014 Oct 17;289(42):28783-94. doi: 10.1074/jbc.M114.597955. Epub 2014 Sep 4. J Biol Chem. 2014. PMID: 25190812 Free PMC article.
-
Structural insights into RNA-dependent eukaryal and archaeal selenocysteine formation.Nucleic Acids Res. 2008 Mar;36(4):1187-99. doi: 10.1093/nar/gkm1122. Epub 2007 Dec 23. Nucleic Acids Res. 2008. PMID: 18158303 Free PMC article.
-
Human SepSecS or SLA/LP: selenocysteine formation and autoimmune hepatitis.Biol Chem. 2010 Jul;391(7):771-6. doi: 10.1515/BC.2010.078. Biol Chem. 2010. PMID: 20623998 Free PMC article. Review.
-
Natural expansion of the genetic code.Nat Chem Biol. 2007 Jan;3(1):29-35. doi: 10.1038/nchembio847. Nat Chem Biol. 2007. PMID: 17173027 Review.
References
-
- Hatfield D.L., Schweizer U., Tsuji P., Gladyshev V.. Selenium: Its Molecular Biology and Role in Human Health. 2016; 4th edn.NY: Springer Science & Business Media.
-
- Schwarz K., Foltz C.M.. Factor 3 activity of selenium compounds. J. Biol. Chem. 1958; 233:245–251. - PubMed
-
- Cox A.G., Tsomides A., Kim A.J., Saunders D., Hwang K.L., Evason K.J., Heidel J., Brown K.K., Yuan M., Lien E.C.et al. .. Selenoprotein H is an essential regulator of redox homeostasis that cooperates with p53 in development and tumorigenesis. Proc. Natl. Acad. Sci. 2016; 113:E5562–E5571. - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

