This is a preprint.
CLCC1 promotes membrane fusion during herpesvirus nuclear egress
- PMID: 39386602
- PMCID: PMC11463520
- DOI: 10.1101/2024.09.23.614151
CLCC1 promotes membrane fusion during herpesvirus nuclear egress
Abstract
Herpesvirales are an ancient viral order that infects species from mollusks to humans for life. During infection, these viruses translocate their large capsids from the nucleus to the cytoplasm independently from the canonical route through the nuclear pore. Instead, capsids dock at the inner nuclear membrane and bud into the perinuclear space. These perinuclear enveloped virions fuse with the outer nuclear membrane releasing the capsids into the cytoplasm for maturation into infectious virions. The budding stage is mediated by virally encoded proteins. But the mediator of the subsequent fusion stage is unknown. Here, using a whole-genome CRISPR screen with herpes simplex virus 1, we identified CLCC1 as an essential host factor for the fusion stage of nuclear egress. Loss of CLCC1 results in a defect in nuclear egress, accumulation of capsid-containing perinuclear vesicles, and a drop in viral titers. In uninfected cells, loss of CLCC1 causes a defect in nuclear pore complex insertion. Viral homologs of CLCC1 are present in herpesviruses that infect mollusks and fish. Our findings uncover an ancient cellular membrane fusion mechanism important for the fundamental cellular process of nuclear envelope morphogenesis that herpesviruses hijack for capsid transport.
Keywords: CLCC1; CRISPR screen; HSV-1; chloride channel; herpes simplex virus; herpesviruses; membrane fusion; nuclear budding; nuclear egress; nuclear pore insertion; nucleocytoplasmic transport.
Figures


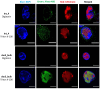










References
-
- Flint S. J., Racaniello V. R., Rall G. F., Hatziioannou T. & Skalka A. M. (Wiley & American Society for Microbiology, Hoboken, NJ, 2020).
-
- Fuchs W., Klupp B. G., Granzow H., Osterrieder N. & Mettenleiter T. C. The interacting UL31 and UL34 gene products of pseudorabies virus are involved in egress from the host-cell nucleus and represent components of primary enveloped but not mature virions. J. Virol. 76, 364–378. (2002). - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
