This is a preprint.
Transient Notch Activation Converts Pluripotent Stem Cell-Derived Cardiomyocytes Towards a Purkinje Fiber Fate
- PMID: 39386729
- PMCID: PMC11463678
- DOI: 10.1101/2024.09.22.614353
Transient Notch Activation Converts Pluripotent Stem Cell-Derived Cardiomyocytes Towards a Purkinje Fiber Fate
Abstract
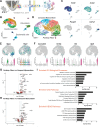
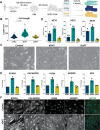
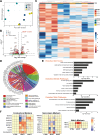
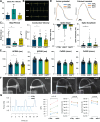
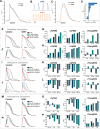
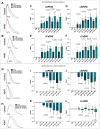
Cardiac Purkinje fibers form the most distal part of the ventricular conduction system. They coordinate contraction and play a key role in ventricular arrhythmias. While many cardiac cell types can be generated from human pluripotent stem cells, methods to generate Purkinje fiber cells remain limited, hampering our understanding of Purkinje fiber biology and conduction system defects. To identify signaling pathways involved in Purkinje fiber formation, we analyzed single cell data from murine embryonic hearts and compared Purkinje fiber cells to trabecular cardiomyocytes. This identified several genes, processes, and signaling pathways putatively involved in cardiac conduction, including Notch signaling. We next tested whether Notch activation could convert human pluripotent stem cell-derived cardiomyocytes to Purkinje fiber cells. Following Notch activation, cardiomyocytes adopted an elongated morphology and displayed altered electrophysiological properties including increases in conduction velocity, spike slope, and action potential duration, all characteristic features of Purkinje fiber cells. RNA-sequencing demonstrated that Notch-activated cardiomyocytes undergo a sequential transcriptome shift, which included upregulation of key Purkinje fiber marker genes involved in fast conduction such as SCN5A, HCN4 and ID2, and downregulation of genes involved in contractile maturation. Correspondingly, we demonstrate that Notch-induced cardiomyocytes have decreased contractile force in bioengineered tissues compared to control cardiomyocytes. We next modified existing in silico models of human pluripotent stem cell-derived cardiomyocytes using our transcriptomic data and modeled the effect of several anti-arrhythmogenic drugs on action potential and calcium transient waveforms. Our models predicted that Purkinje fiber cells respond more strongly to dofetilide and amiodarone, while cardiomyocytes are more sensitive to treatment with nifedipine. We validated these findings in vitro, demonstrating that our new cell-specific in vitro model can be utilized to better understand human Purkinje fiber physiology and its relevance to disease.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures

References
-
- Árpádffy-Lovas T., Husti Z., Baczkó I., Varró A. and Virág L. (2021). Different effects of amiodarone and dofetilide on the dispersion of repolarization between well-coupled ventricular and Purkinje fibers1. Can. J. Physiol. Pharmacol. 99, 48–55. - PubMed
-
- Baruteau A.-E., Probst V. and Abriel H. (2015). Inherited progressive cardiac conduction disorders. Curr Opin Cardiol 30, 33–39. - PubMed
-
- Bray N. L., Pimentel H., Melsted P. and Pachter L. (2016). Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol 34, 525–527. - PubMed
-
- Bruneau B. G. and Srivastava D. (2014). Congenital Heart Disease. Circulation Research 114, 598–599. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
