Uncovering lipid dynamics in Staphylococcus aureus osteomyelitis using multimodal imaging mass spectrometry
- PMID: 39389064
- PMCID: PMC11977171
- DOI: 10.1016/j.chembiol.2024.09.005
Uncovering lipid dynamics in Staphylococcus aureus osteomyelitis using multimodal imaging mass spectrometry
Abstract
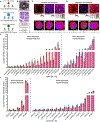
Osteomyelitis occurs when Staphylococcus aureus invades the bone microenvironment, resulting in a bone marrow abscess with a spatially defined architecture of cells and biomolecules. Imaging mass spectrometry and microscopy are tools that can be employed to interrogate the lipidome of S. aureus-infected murine femurs and reveal metabolic and signaling consequences of infection. Here, nearly 250 lipids were spatially mapped to healthy and infection-associated morphological features throughout the femur, establishing composition profiles for tissue types. Ether lipids and arachidonoyl lipids were altered between cells and tissue structures in abscesses, suggesting their roles in abscess formation and inflammatory signaling. Sterols, triglycerides, bis(monoacylglycero)phosphates, and gangliosides possessed ring-like distributions throughout the abscess, suggesting a hypothesized dysregulation of lipid metabolism in a population of cells that cannot be discerned with traditional microscopy. These data provide insight into the signaling function and metabolism of cells in the fibrotic border of abscesses, likely characteristic of lipid-laden macrophages.
Keywords: Imging mass spectrometry; arachidonic acid; ether lipids; foam cells; lipidomics; macrophages; osteomyelitis; spatial biology; staphylococcus aureus.
Copyright © 2024 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures
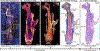





Update of
-
In situ lipidomics of Staphylococcus aureus osteomyelitis using imaging mass spectrometry.bioRxiv [Preprint]. 2023 Dec 2:2023.12.01.569690. doi: 10.1101/2023.12.01.569690. bioRxiv. 2023. Update in: Cell Chem Biol. 2024 Oct 17;31(10):1852-1868.e5. doi: 10.1016/j.chembiol.2024.09.005. PMID: 38077019 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

