A sphingolipid rheostat controls apoptosis versus apical cell extrusion as alternative tumour-suppressive mechanisms
- PMID: 39397024
- PMCID: PMC11471799
- DOI: 10.1038/s41419-024-07134-2
A sphingolipid rheostat controls apoptosis versus apical cell extrusion as alternative tumour-suppressive mechanisms
Abstract
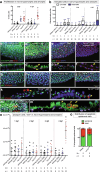
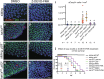
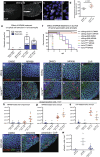
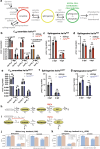
Evasion of cell death is a hallmark of cancer, and consequently the induction of cell death is a common strategy in cancer treatment. However, the molecular mechanisms regulating different types of cell death are poorly understood. We have formerly shown that in the epidermis of hypomorphic zebrafish hai1a mutant embryos, pre-neoplastic transformations of keratinocytes caused by unrestrained activity of the type II transmembrane serine protease Matriptase-1 heal spontaneously. This healing is driven by Matriptase-dependent increased sphingosine kinase (SphK) activity and sphingosine-1-phosphate (S1P)-mediated keratinocyte loss via apical cell extrusion. In contrast, amorphic hai1afr26 mutants with even higher Matriptase-1 and SphK activity die within a few days. Here we show that this lethality is not due to epidermal carcinogenesis, but to aberrant tp53-independent apoptosis of keratinocytes caused by increased levels of pro-apoptotic C16 ceramides, sphingolipid counterparts to S1P within the sphingolipid rheostat, which severely compromises the epidermal barrier. Mathematical modelling of sphingolipid rheostat homeostasis, combined with in vivo manipulations of components of the rheostat or the ceramide de novo synthesis pathway, indicate that this unexpected overproduction of ceramides is caused by a negative feedback loop sensing ceramide levels and controlling ceramide replenishment via de novo synthesis. Therefore, despite their initial decrease due to increased conversion to S1P, ceramides eventually reach cell death-inducing levels, making transformed pre-neoplastic keratinocytes die even before they are extruded, thereby abrogating the normally barrier-preserving mode of apical live cell extrusion. Our results offer an in vivo perspective of the dynamics of sphingolipid homeostasis and its relevance for epithelial cell survival versus cell death, linking apical cell extrusion and apoptosis. Implications for human carcinomas and their treatments are discussed.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


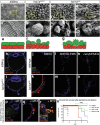
References
MeSH terms
Substances
Grants and funding
- FOR 5151/Deutsche Forschungsgemeinschaft (German Research Foundation)
- SFB 829/Deutsche Forschungsgemeinschaft (German Research Foundation)
- 453407124/Deutsche Forschungsgemeinschaft (German Research Foundation)
- EXC 2075 - 390740016/Deutsche Forschungsgemeinschaft (German Research Foundation)
- R01 GM063904/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

