LETMD1 regulates mitochondrial protein synthesis and import to guard brown fat mitochondrial integrity and function
- PMID: 39398236
- PMCID: PMC11467678
- DOI: 10.1016/j.isci.2024.110944
LETMD1 regulates mitochondrial protein synthesis and import to guard brown fat mitochondrial integrity and function
Abstract
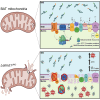
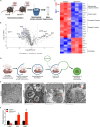
Thermogenic brown adipocytes (BAs) catabolize lipids to generate heat, representing powerful agents against the growing global obesity epidemic. We and others reported recently that LETMD1 is a BA-specific protein essential for mitochondrial structure and function, but the mechanisms of action remain unclear. We performed sequential digestion to demonstrate that LETMD1 is a trans-inner mitochondrial membrane protein. We then generated UCP1Cre-driven BA-specific Letmd1 knockout (Letmd1 UKO ) mice to show that Letmd1 UKO leads to protein aggregation, reactive oxidative stress, hyperpolarization, and mitophagy in BAs. We further employed TurboID proximity labeling to identify LETMD1-interacting proteins. Many candidate proteins are associated with mitochondrial ribosomes, protein import machinery, and electron transport chain complexes (ETC-I and ETC-IV). Using quantitative proteomics, we confirmed the elevated aggregations of ETC and mitochondrial ribosomal proteins, impairing mitochondrial protein synthesis in the Letmd1 UKO BAs. Therefore, LETMD1 may function to maintain mitochondrial proteostasis through regulating import of nuclear-encoded proteins and local protein translation in brown fat mitochondria.
Keywords: Cell; Cell biology; Cellular physiology; Molecular biology.
© 2024 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

