An efficient CRISPR-Cas12a-mediated MicroRNA knockout strategy in plants
- PMID: 39401095
- PMCID: PMC11672732
- DOI: 10.1111/pbi.14484
An efficient CRISPR-Cas12a-mediated MicroRNA knockout strategy in plants
Abstract
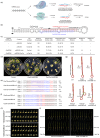
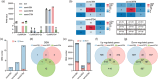
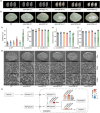
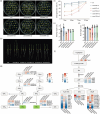
In recent years, the CRISPR-Cas9 nuclease has been used to knock out MicroRNA (miRNA) genes in plants, greatly promoting the study of miRNA function. However, due to its propensity for generating small insertions and deletions, Cas9 is not well-suited for achieving a complete knockout of miRNA genes. By contrast, CRISPR-Cas12a nuclease generates larger deletions, which could significantly disrupt the secondary structure of pre-miRNA and prevent the production of mature miRNAs. Through the case study of OsMIR390 in rice, we confirmed that Cas12a is a more efficient tool than Cas9 in generating knockout mutants of a miRNA gene. To further demonstrate CRISPR-Cas12a-mediated knockout of miRNA genes in rice, we targeted nine OsMIRNA genes that have different spaciotemporal expression and have not been previously investigated via genetic knockout approaches. With CRISPR-Cas12a, up to 100% genome editing efficiency was observed at these miRNA loci. The resulting larger deletions suggest Cas12a robustly generated null alleles of miRNA genes. Transcriptome profiling of the miRNA mutants, as well as phenotypic analysis of the rice grains revealed the function of these miRNAs in controlling gene expression and regulating grain quality and seed development. This study established CRISPR-Cas12a as an efficient tool for genetic knockout of miRNA genes in plants.
Keywords: CRISPR‐Cas12a; MicroRNA; genome Editing; germplasm innovation; rice.
© 2024 The Author(s). Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
-
- Bernardi, Y. , Ponso, M.A. , Belen, F. , Vegetti, A.C. and Dotto, M.C. (2022) MicroRNA miR394 regulates flowering time in Arabidopsis thaliana. Plant Cell Rep. 41, 1375–1388. - PubMed
-
- Cheng, X. , He, Q. , Tang, S. , Wang, H. , Zhang, X. , Lv, M. , Liu, H. et al. (2021) The miR172/IDS1 signaling module confers salt tolerance through maintaining ROS homeostasis in cereal crops. New Phytol. 230, 1017–1033. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

