Pharmacology of PIEZO1 channels
- PMID: 39402010
- PMCID: PMC7618196
- DOI: 10.1111/bph.17351
Pharmacology of PIEZO1 channels
Abstract
PIEZO1 is a eukaryotic membrane protein that assembles as trimers to form calcium-permeable, non-selective cation channels with exquisite capabilities for mechanical force sensing and transduction of force into effect in diverse cell types that include blood cells, endothelial cells, epithelial cells, fibroblasts and stem cells and diverse systems that include bone, lymphatics and muscle. The channel has wide-ranging roles and is considered as a target for novel therapeutics in ailments spanning cancers and cardiovascular, dental, gastrointestinal, hepatobiliary, infectious, musculoskeletal, nervous system, ocular, pregnancy, renal, respiratory and urological disorders. The identification of PIEZO1 modulators is in its infancy but useful experimental tools emerged for activating, and to a lesser extent inhibiting, the channels. Elementary structure-activity relationships are known for the Yoda series of small molecule agonists, which show the potential for diverse physicochemical and pharmacological properties. Intriguing effects of Yoda1 include the stimulated removal of excess cerebrospinal fluid. Despite PIEZO1's broad expression, opportunities are suggested for selective positive or negative modulation without intolerable adverse effects. Here we provide a focused, non-systematic, narrative review of progress with this pharmacology and discuss potential future directions for research in the area.
Keywords: calcium ion; ion channel; mechanical force; non‐selective cation channel; pharmacology; pressure; shear stress; small molecule modulator.
© 2024 The Author(s). British Journal of Pharmacology published by John Wiley & Sons Ltd on behalf of British Pharmacological Society.
Conflict of interest statement
D. J. B. and R. F. are partners of CalTIC GmbH, a pharmaceutical startup company with a mission to develop ion channel modulators as new classes of medicines. No other conflicts of interests are disclosed.
Figures
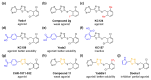
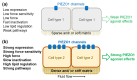

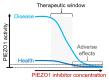


References
-
- Albuisson J, Murthy SE, Bandell M, Coste B, Louis-Dit-Picard H, Mathur J, Fénéant-Thibault M, Tertian G, de Jaureguiberry JP, Syfuss PY, Cahalan S. Dehydrated hereditary stomatocytosis linked to gain-of-function mutations in mechanically activated PIEZO1 ion channels. Nature Communications. 2013;4:1884. doi: 10.1038/ncomms2899. - DOI - PMC - PubMed
-
- Alexander SPH, Christopoulos A, Davenport AP, Kelly E, Mathie AA, Peters JA, Veale EL, Armstrong JF, Faccenda E, Harding SD, Davies JA, et al. The Concise Guide to PHARMACOLOGY 2023/24: G protein-coupled receptors. British Journal of Pharmacology. 2023;180(Suppl 2):S23–S144. - PubMed
-
- Alexander SPH, Mathie AA, Peters JA, Veale EL, Striessnig J, Kelly E, Armstrong JF, Faccenda E, Harding SD, Davies JA, Aldrich RW, et al. The concise guide to PHARMACOLOGY 2023/24: Ion channels. British Journal of Pharmacology. 2023;180(Suppl 2):S145–S222. doi: 10.1111/bph.16178. - DOI - PMC - PubMed
-
- Amado NG, Nosyreva ED, Thompson D, Egeland TJ, Ogujiofor OW, Yang M, Fusco AN, Passoni N, Mathews J, Cantarel B, Baker LA, et al. PIEZO1 loss-of-function compound heterozygous mutations in the rare congenital human disorder prune belly syndrome. Nature Communications. 2024;15:339. doi: 10.1038/s41467-023-44594-0. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

