Elevated Aβ aggregates in feces from Alzheimer's disease patients: a proof-of-concept study
- PMID: 39402637
- PMCID: PMC11472473
- DOI: 10.1186/s13195-024-01597-3
Elevated Aβ aggregates in feces from Alzheimer's disease patients: a proof-of-concept study
Abstract
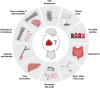
Background: Misfolding and aggregation of amyloid β (Aβ), along with neurofibrillary tangles consisting of aggregated Tau species, are pathological hallmarks of Alzheimer's disease (AD) onset and progression. In this study, we hypothesized the clearance of Aβ aggregates from the brain and body into the gut.
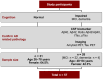
Methods: To investigate this, we used surface-based fluorescence intensity distribution analysis (sFIDA) to determine the Aβ aggregate concentrations in feces from 26 AD patients and 31 healthy controls (HC).
Results: Aβ aggregates were detectable in human feces and their concentrations were elevated in AD patients compared to HC (specificity 90.3%, sensitivity 53.8%).
Conclusion: Thus, fecal Aβ aggregates constitute a non-invasive biomarker candidate for diagnosing AD. Whether digestion-resistant Aβ aggregates in feces are secreted via the liver and bile or directly from the enteric neuronal system remains to be elucidated.
Keywords: Amyloidosis; Aβ oligomer quantitation; Brain-gut-microbiota axis; Clearance; Fecal/stool samples; Leaky gut; sFIDA.
© 2024. The Author(s).
Conflict of interest statement
D.W. and O.B. are cofounders and shareholders of attyloid GmbH. D.W. is member of attyloid’s supervisory board. M.P., O.B., G.T. and D.W. are inventors of patent DE102020114278A1 (determination of disease-specific protein aggregates in fecal samples). These factors did not influence the interpretation of the data. Remaining authors declare no conflicts of interest.
Figures




References
-
- Kulenkampff K, Wolf Perez AM, Sormanni P, Habchi J, Vendruscolo M. Quantifying misfolded protein oligomers as drug targets and biomarkers in Alzheimer and Parkinson diseases. Nat Reviews Chem. 2021;5(4):277–94. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

