Androgen signaling restricts glutaminolysis to drive sex-specific Th17 metabolism in allergic airway inflammation
- PMID: 39404231
- PMCID: PMC11601904
- DOI: 10.1172/JCI177242
Androgen signaling restricts glutaminolysis to drive sex-specific Th17 metabolism in allergic airway inflammation
Abstract
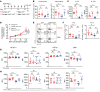
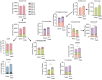
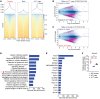
Female individuals have an increased prevalence of many Th17 cell-mediated diseases, including asthma. Androgen signaling decreases Th17 cell-mediated airway inflammation, and Th17 cells rely on glutaminolysis. However, it remains unclear whether androgen receptor (AR) signaling modifies glutamine metabolism to suppress Th17 cell-mediated airway inflammation. We show that Th17 cells from male humans and mice had decreased glutaminolysis compared with female individuals, and that AR signaling attenuated Th17 cell mitochondrial respiration and glutaminolysis in mice. Using allergen-induced airway inflammation mouse models, we determined that females had a selective reliance upon glutaminolysis for Th17-mediated airway inflammation, and that AR signaling attenuated glutamine uptake in CD4+ T cells by reducing expression of glutamine transporters. In patients with asthma, circulating Th17 cells from men had minimal reliance upon glutamine uptake compared to Th17 cells from women. AR signaling thus attenuates glutaminolysis, demonstrating sex-specific metabolic regulation of Th17 cells with implications for Th17 or glutaminolysis targeted therapeutics.
Keywords: Asthma; Immunology; Pulmonology; Sex hormones; T cells.
Conflict of interest statement
Figures





Comment in
- Sex, cells, and metabolism: Androgens temper Th17-mediated immunity doi: 10.1172/JCI186520
References
MeSH terms
Substances
Grants and funding
- U01 AI155299/AI/NIAID NIH HHS/United States
- R01 HL122554/HL/NHLBI NIH HHS/United States
- T32 AI138932/AI/NIAID NIH HHS/United States
- F30 HL159941/HL/NHLBI NIH HHS/United States
- T32 GM139800/GM/NIGMS NIH HHS/United States
- T32 GM007347/GM/NIGMS NIH HHS/United States
- I01 BX004299/BX/BLRD VA/United States
- F31 CA261049/CA/NCI NIH HHS/United States
- R01 HL136664/HL/NHLBI NIH HHS/United States
- P30 DK058404/DK/NIDDK NIH HHS/United States
- K00 CA253718/CA/NCI NIH HHS/United States
- T32 DK101003/DK/NIDDK NIH HHS/United States
- U19 AI162310/AI/NIAID NIH HHS/United States
- T32 HL094296/HL/NHLBI NIH HHS/United States
- F30 CA239367/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

