Tissue-resident memory T cells contribute to protection against heterologous SARS-CoV-2 challenge
- PMID: 39405115
- PMCID: PMC11623939
- DOI: 10.1172/jci.insight.184074
Tissue-resident memory T cells contribute to protection against heterologous SARS-CoV-2 challenge
Abstract
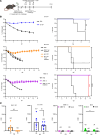
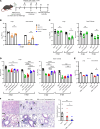
New vaccine formulations are based on circulating strains of virus, which have tended to evolve to more readily transmit human to human and to evade the neutralizing antibody response. An assumption of this approach is that ancestral strains of virus will not recur. Recurrence of these strains could be a problem for individuals not previously exposed to ancestral spike protein. Here, we addressed this by infecting mice with recent SARS-CoV-2 variants and then challenging them with a highly pathogenic mouse-adapted virus closely related to the ancestral Wuhan-1 strain (SARS2-N501YMA30). We found that challenged mice were protected from severe disease, despite having low or no neutralizing antibodies against SARS2-N501YMA30. T cell depletion from previously infected mice did not diminish infection against clinical disease, although it resulted in delayed virus clearance in the nasal turbinate and, in some cases, in the lungs. Levels of tissue-resident memory T cells were significantly elevated in the nasal turbinate of previously infected mice compared with that of naive mice. However, this phenotype was not seen in lung tissues. Together, these results indicate that the immune response to newly circulating variants afforded protection against reinfection with the ancestral virus that was in part T cell based.
Keywords: Adaptive immunity; COVID-19; T cells; Virology.
Conflict of interest statement
Figures



References
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

