Multiomics profiling of mouse polycystic kidney disease progression at a single-cell resolution
- PMID: 39405347
- PMCID: PMC11513963
- DOI: 10.1073/pnas.2410830121
Multiomics profiling of mouse polycystic kidney disease progression at a single-cell resolution
Abstract
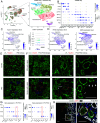
Autosomal dominant polycystic kidney disease (ADPKD) is the most common hereditary kidney disease and causes significant morbidity, ultimately leading to kidney failure. PKD pathogenesis is characterized by complex and dynamic alterations in multiple cell types during disease progression, hampering a deeper understanding of disease mechanism and the development of therapeutic approaches. Here, we generate a single-nucleus multimodal atlas of an orthologous mouse PKD model at early, mid, and late timepoints, consisting of 125,434 single-nucleus transcriptomic and epigenetic multiomes. We catalog differentially expressed genes and activated epigenetic regions in each cell type during PKD progression, characterizing cell-type-specific responses to Pkd1 deletion. We describe heterogeneous, atypical collecting duct cells as well as proximal tubular cells that constitute cyst epithelia in PKD. The transcriptional regulation of the cyst lining cell marker GPRC5A is conserved between mouse and human PKD cystic epithelia, suggesting shared gene regulatory pathways. Our single-nucleus multiomic analysis of mouse PKD provides a foundation to understand the earliest changes molecular deregulation in a mouse model of PKD at a single-cell resolution.
Keywords: PKD1; mouse model; multiomics; polycystic kidney disease; single cell analysis.
Conflict of interest statement
Competing interests statement:B.D.H. is a consultant for Janssen Research & Development, LLC, Pfizer and Chinook Therapeutics. B.D.H holds equity in Chinook Therapeutics. B.D.H holds grant funding from Chinook Therapeutics and Janssen Research & Development, LLC. O.M.W has received grants from AstraZeneca unrelated to the current work.
Figures




Update of
-
Multi-omics profiling of mouse polycystic kidney disease progression at a single cell resolution.bioRxiv [Preprint]. 2024 May 31:2024.05.27.595830. doi: 10.1101/2024.05.27.595830. bioRxiv. 2024. Update in: Proc Natl Acad Sci U S A. 2024 Oct 22;121(43):e2410830121. doi: 10.1073/pnas.2410830121. PMID: 38854144 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
- R01 DK103740/DK/NIDDK NIH HHS/United States
- 2R01DK103740/HHS | NIH | National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK)
- UC2DK126024/HHS | NIH | National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK)
- UC2 DK126024/DK/NIDDK NIH HHS/United States
- U24 DK126110/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

