Insights into the Molecular Structure, Stability, and Biological Significance of Non-Canonical DNA Forms, with a Focus on G-Quadruplexes and i-Motifs
- PMID: 39407611
- PMCID: PMC11477922
- DOI: 10.3390/molecules29194683
Insights into the Molecular Structure, Stability, and Biological Significance of Non-Canonical DNA Forms, with a Focus on G-Quadruplexes and i-Motifs
Abstract
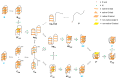
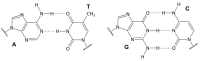
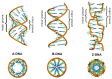
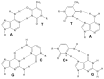
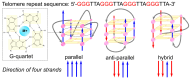
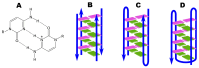
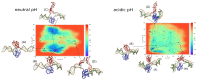
This article provides a comprehensive examination of non-canonical DNA structures, particularly focusing on G-quadruplexes (G4s) and i-motifs. G-quadruplexes, four-stranded structures formed by guanine-rich sequences, are stabilized by Hoogsteen hydrogen bonds and monovalent cations like potassium. These structures exhibit diverse topologies and are implicated in critical genomic regions such as telomeres and promoter regions of oncogenes, playing significant roles in gene expression regulation, genome stability, and cellular aging. I-motifs, formed by cytosine-rich sequences under acidic conditions and stabilized by hemiprotonated cytosine-cytosine (C:C+) base pairs, also contribute to gene regulation despite being less prevalent than G4s. This review highlights the factors influencing the stability and dynamics of these structures, including sequence composition, ionic conditions, and environmental pH. Molecular dynamics simulations and high-resolution structural techniques have been pivotal in advancing our understanding of their folding and unfolding mechanisms. Additionally, the article discusses the therapeutic potential of small molecules designed to selectively bind and stabilize G4s and i-motifs, with promising implications for cancer treatment. Furthermore, the structural properties of these DNA forms are explored for applications in nanotechnology and molecular devices. Despite significant progress, challenges remain in observing these structures in vivo and fully elucidating their biological functions. The review underscores the importance of continued research to uncover new insights into the genomic roles of G4s and i-motifs and their potential applications in medicine and technology. This ongoing research promises exciting developments in both basic science and applied fields, emphasizing the relevance and future prospects of these intriguing DNA structures.
Keywords: G-quadruplex; i-motif; ligands; molecular dynamics; multi-stranded DNA structures; stability.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


Similar articles
-
DNA G-Quadruplex in Human Telomeres and Oncogene Promoters: Structures, Functions, and Small Molecule Targeting.Acc Chem Res. 2022 Sep 20;55(18):2628-2646. doi: 10.1021/acs.accounts.2c00337. Epub 2022 Sep 2. Acc Chem Res. 2022. PMID: 36054116 Free PMC article.
-
G-quadruplex DNA: A Longer Story.Acc Chem Res. 2022 Nov 15;55(22):3242-3252. doi: 10.1021/acs.accounts.2c00519. Epub 2022 Oct 25. Acc Chem Res. 2022. PMID: 36282946
-
Structural diversity and specific recognition of four stranded G-quadruplex DNA.Curr Mol Med. 2011 Dec;11(9):744-69. doi: 10.2174/156652411798062421. Curr Mol Med. 2011. PMID: 21999150 Review.
-
An Updated Focus on Quadruplex Structures as Potential Therapeutic Targets in Cancer.Int J Mol Sci. 2020 Nov 24;21(23):8900. doi: 10.3390/ijms21238900. Int J Mol Sci. 2020. PMID: 33255335 Free PMC article. Review.
-
Structurally diverse G-quadruplexes as the noncanonical nucleic acid drug target for live cell imaging and antibacterial study.Chem Commun (Camb). 2023 Feb 2;59(11):1415-1433. doi: 10.1039/d2cc05945b. Chem Commun (Camb). 2023. PMID: 36636928 Review.
Cited by
-
Polymeric Properties of Telomeric G-Quadruplex Multimers: Effects of Chemically Inert Crowders.Biomacromolecules. 2025 May 12;26(5):3128-3138. doi: 10.1021/acs.biomac.5c00176. Epub 2025 Apr 8. Biomacromolecules. 2025. PMID: 40199738 Free PMC article.
-
RNA G-quadruplexes: emerging regulators of gene expression and therapeutic targets.Funct Integr Genomics. 2025 Jul 3;25(1):143. doi: 10.1007/s10142-025-01656-4. Funct Integr Genomics. 2025. PMID: 40608121 Review.
-
Conformational Propensities of a DNA Hairpin with a Stem Sequence from the c-MYC Promoter.Biomolecules. 2025 Mar 26;15(4):483. doi: 10.3390/biom15040483. Biomolecules. 2025. PMID: 40305258 Free PMC article.
-
G-Quadruplex-Based Splice Switching as a Therapeutic Approach in Duchenne Muscular Dystrophy.ACS Chem Biol. 2025 Mar 21;20(3):670-679. doi: 10.1021/acschembio.4c00805. Epub 2025 Mar 3. ACS Chem Biol. 2025. PMID: 40029284 Free PMC article.
References
-
- Berg J.M., Tymoczko J.L., Gatto G.J., Stryer L. Biochemistry. 9th ed. Macmillan International, Higher Education; New York, NY, USA: 2019.
-
- Allison L.A. Fundamental Molecular Biology. 3rd ed. Wiley Blackwell; Hoboken, NJ, USA: Chichester, UK: 2021.
-
- Neidle S., editor. Oxford Handbook of Nucleic Acid Structure. Oxford University Press; Oxford, UK: New York, NY, USA: 1999.
-
- Turner P.C., editor. Molecular Biology. 3rd ed. Taylor & Francis; New York, NY, USA: 2005. BIOS instant notes.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

