DNA Origami Incorporated into Solid-State Nanopores Enables Enhanced Sensitivity for Precise Analysis of Protein Translocations
- PMID: 39420456
- PMCID: PMC12266972
- DOI: 10.1021/acs.analchem.4c02016
DNA Origami Incorporated into Solid-State Nanopores Enables Enhanced Sensitivity for Precise Analysis of Protein Translocations
Abstract
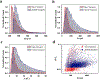
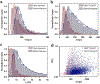
The rapidly advancing field of nanotechnology is driving the development of precise sensing methods at the nanoscale, with solid-state nanopores emerging as promising tools for biomolecular sensing. This study investigates the increased sensitivity of solid-state nanopores achieved by integrating DNA origami structures, leading to the improved analysis of protein translocations. Using holo human serum transferrin (holo-hSTf) as a model protein, we compared hybrid nanopores incorporating DNA origami with open solid-state nanopores. Results show a significant enhancement in holo-hSTf detection sensitivity with DNA origami integration, suggesting a unique role of DNA interactions beyond confinement. This approach holds potential for ultrasensitive protein detection in biosensing applications, offering advancements in biomedical research and diagnostic tool development for diseases with low-abundance protein biomarkers. Further exploration of origami designs and nanopore configurations promises even greater sensitivity and versatility in the detection of a wider range of proteins, paving the way for advanced biosensing technologies.
Figures



Similar articles
-
Aerolysin Nanopore Electrochemistry.Acc Chem Res. 2025 Feb 18;58(4):517-528. doi: 10.1021/acs.accounts.4c00630. Epub 2025 Jan 28. Acc Chem Res. 2025. PMID: 39874057 Review.
-
The Potential of Nanopore Technologies in Peptide and Protein Sensing for Biomarker Detection.Biosensors (Basel). 2025 Aug 16;15(8):540. doi: 10.3390/bios15080540. Biosensors (Basel). 2025. PMID: 40863000 Free PMC article. Review.
-
Supercharged DNA origami enhanced signal amplification for ultrasensitive detection of nucleic acid.Biosens Bioelectron. 2025 Nov 1;287:117692. doi: 10.1016/j.bios.2025.117692. Epub 2025 Jun 13. Biosens Bioelectron. 2025. PMID: 40517661
-
Site-Specific Integration of Hexagonal Boron Nitride Quantum Emitters on 2D DNA Origami Nanopores.Nano Lett. 2024 Jul 17;24(28):8510-8517. doi: 10.1021/acs.nanolett.4c00673. Epub 2024 Jun 10. Nano Lett. 2024. PMID: 38856705 Free PMC article.
-
Specific ATP Detection Using Molecule-Responsive DNA Nanopores.Small. 2025 Jul;21(29):e2409293. doi: 10.1002/smll.202409293. Epub 2025 May 2. Small. 2025. PMID: 40317998 Free PMC article.
Cited by
-
Solid-State Nanopore Analysis of DNA-Templated Silver Nanoclusters: Voltage-Dependent Translocation and Electrolyte Stability.ACS Appl Mater Interfaces. 2025 Jul 9;17(27):39407-39419. doi: 10.1021/acsami.5c02405. Epub 2025 May 27. ACS Appl Mater Interfaces. 2025. PMID: 40420685 Free PMC article.
-
Engineering DNA nanopores: from structural evolution to sensing and transport.Mater Today Bio. 2025 Jul 26;34:102137. doi: 10.1016/j.mtbio.2025.102137. eCollection 2025 Oct. Mater Today Bio. 2025. PMID: 40761509 Free PMC article. Review.
References
-
- Chen Z; Wang C; Hao L; Gao R; Li F; Liu S Biosens. Bioelectron 2019, 128, 104–112. - PubMed
-
- Meunier-Prest R; Bouyon A; Rampazzi E; Raveau S; Andreoletti P; Cherkaoui-Malki M Biosens. Bioelectron 2010, 25 (12), 2598–2602. - PubMed
-
- Diculescu VC; Chiorcea-Paquim A-M; Oliveira-Brett AM TrAC, Trends Anal. Chem 2016, 79, 23–36.
-
- Song S; Wang L; Li J; Fan C; Zhao J TrAC, Trends Anal. Chem 2008, 27 (2), 108–117.
-
- Vierstra J; Stamatoyannopoulos JA Nat. Methods 2016, 13 (3), 213–221. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

