Mutagenicity and genotoxicity evaluation of 15 nitrosamine drug substance-related impurities in human TK6 cells
- PMID: 39433234
- PMCID: PMC12477663
- DOI: 10.1016/j.yrtph.2024.105730
Mutagenicity and genotoxicity evaluation of 15 nitrosamine drug substance-related impurities in human TK6 cells
Abstract
Nitrosamine drug substance-related impurities (NDSRIs) are a sub-category of N-nitrosamine drug impurities that share structural similarity to the corresponding active pharmaceutical ingredient. The mutagenicity of NDSRIs is poorly understood. We previously tested a series of NDSRIs using the Enhanced Ames Test (EAT). In this follow-up study, we further examined the genotoxicity and mutagenicity of 15 of these NDSRIs in human TK6 cells. Seven EAT-positive NDSRIs, including N-nitroso-nortriptyline, N-nitroso-fluoxetine, N-nitroso-desmethyl-diphenhydramine, N-nitroso-duloxetine, N-nitroso-lorcaserin, N-nitroso-varenicline, and N-nitroso-sertraline, induced concentration-dependent increases in micronuclei after bioactivation with hamster liver S9. These NDSRIs were also mutagenic in the TK and HPRT gene mutation assays, consistent with their positive EAT results. In the presence of hamster liver S9, the eight EAT-negative NDSRIs were negative in the micronucleus assay and negative for mutation induction. Using TK6 cells endogenously expressing a single human cytochrome P450 (CYP), we found that CYP2C19, CYP2B6, CYP2A6, and CYP3A4 are key enzymes activating the genotoxicity and mutagenicity of these NDSRIs. Overall, the hamster S9-mediated TK6 cell mutagenicity results agreed with those observed in the EAT, indicating consistency in the mutagenic responses produced by NDSRIs across different testing systems. These data support the use of EAT for hazard identification and safety assessment of NDSRIs.
Keywords: Chromosomal damage; DNA damage; Gene mutation; Hamster liver S9; NDSRIs; Nitrosamine impurities.
Copyright © 2024. Published by Elsevier Inc.
Conflict of interest statement
Declaration of competing interest This article reflects the views of the authors and does not necessarily reflect those of the U.S. Food and Drug Administration (FDA). Any mention of commercial products is for clarification only and is not intended as approval, endorsement, or recommendation. The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

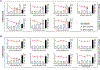



References
-
- Ando M, et al. , 2014. Usefulness of monitoring γ-H2AX and cell cycle arrest in HepG2 cells for estimating genotoxicity using a high-content analysis system. J. Biomol. Screen 19, 1246–1254. - PubMed
-
- Bellec G, et al. , 1996. Cytochrome P450 metabolic dealkylation of nine N-nitrosodialkylamines by human liver microsomes. Carcinogenesis 17, 2029–2034. - PubMed
-
- Bharate SS, 2021. Critical analysis of drug product recalls due to nitrosamine impurities. J. Med. Chem. 64, 2923–2936. - PubMed
-
- Brambilla G, et al. , 1985. Formation of DNA-damaging nitroso compounds by interaction of drugs with nitrite. A preliminary screening for detecting potentially hazardous drugs. J. Toxicol. Environ. Health 15, 1–24. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

