Functional analysis of cyclic diguanylate-modulating proteins in Vibrio fischeri
- PMID: 39436151
- PMCID: PMC11575326
- DOI: 10.1128/msystems.00956-24
Functional analysis of cyclic diguanylate-modulating proteins in Vibrio fischeri
Abstract
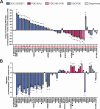
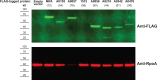
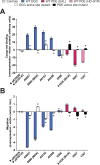
As bacterial symbionts transition from a motile free-living state to a sessile biofilm state, they must coordinate behavior changes suitable to each lifestyle. Cyclic diguanylate (c-di-GMP) is an intracellular signaling molecule that can regulate this transition, and it is synthesized by diguanylate cyclase (DGC) enzymes and degraded by phosphodiesterase (PDE) enzymes. Generally, c-di-GMP inhibits motility and promotes biofilm formation. While c-di-GMP and the enzymes that contribute to its metabolism have been well studied in pathogens, considerably less focus has been placed on c-di-GMP regulation in beneficial symbionts. Vibrio fischeri is the sole beneficial symbiont of the Hawaiian bobtail squid (Euprymna scolopes) light organ, and the bacterium requires both motility and biofilm formation to efficiently colonize. c-di-GMP regulates swimming motility and cellulose exopolysaccharide production in V. fischeri. The genome encodes 50 DGCs and PDEs, and while a few of these proteins have been characterized, the majority have not undergone comprehensive characterization. In this study, we use protein overexpression to systematically characterize the functional potential of all 50 V. fischeri proteins. All 28 predicted DGCs and 10 of the 14 predicted PDEs displayed at least one phenotype consistent with their predicted function, and a majority of each displayed multiple phenotypes. Finally, active site mutant analysis of proteins with the potential for both DGC and PDE activities revealed potential activities for these proteins. This work presents a systems-level functional analysis of a family of signaling proteins in a tractable animal symbiont and will inform future efforts to characterize the roles of individual proteins during lifestyle transitions.IMPORTANCECyclic diguanylate (c-di-GMP) is a critical second messenger that mediates bacterial behaviors, and Vibrio fischeri colonization of its Hawaiian bobtail squid host presents a tractable model in which to interrogate the role of c-di-GMP during animal colonization. This work provides systems-level characterization of the 50 proteins predicted to modulate c-di-GMP levels. By combining multiple assays, we generated a rich understanding of which proteins have the capacity to influence c-di-GMP levels and behaviors. Our functional approach yielded insights into how proteins with domains to both synthesize and degrade c-di-GMP may impact bacterial behaviors. Finally, we integrated published data to provide a broader picture of each of the 50 proteins analyzed. This study will inform future work to define specific pathways by which c-di-GMP regulates symbiotic behaviors and transitions.
Keywords: Vibrio fischeri; biofilm; c-di-GMP; flagellar motility.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Update of
-
Functional analysis of cyclic diguanylate-modulating proteins in Vibrio fischeri.bioRxiv [Preprint]. 2024 Sep 24:2023.07.24.550417. doi: 10.1101/2023.07.24.550417. bioRxiv. 2024. Update in: mSystems. 2024 Nov 19;9(11):e0095624. doi: 10.1128/msystems.00956-24. PMID: 37546929 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

