Binding profiles for 961 Drosophila and C. elegans transcription factors reveal tissue-specific regulatory relationships
- PMID: 39438113
- PMCID: PMC11694743
- DOI: 10.1101/gr.279037.124
Binding profiles for 961 Drosophila and C. elegans transcription factors reveal tissue-specific regulatory relationships
Abstract
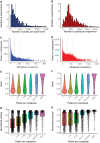
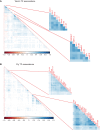
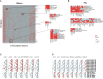
A catalog of transcription factor (TF) binding sites in the genome is critical for deciphering regulatory relationships. Here, we present the culmination of the efforts of the modENCODE (model organism Encyclopedia of DNA Elements) and modERN (model organism Encyclopedia of Regulatory Networks) consortia to systematically assay TF binding events in vivo in two major model organisms, Drosophila melanogaster (fly) and Caenorhabditis elegans (worm). These data sets comprise 605 TFs identifying 3.6 M sites in the fly and 356 TFs identifying 0.9 M sites in the worm, and represent the majority of the regulatory space in each genome. We demonstrate that TFs associate with chromatin in clusters termed "metapeaks," that larger metapeaks have characteristics of high-occupancy target (HOT) regions, and that the importance of consensus sequence motifs bound by TFs depends on metapeak size and complexity. Combining ChIP-seq data with single-cell RNA-seq data in a machine-learning model identifies TFs with a prominent role in promoting target gene expression in specific cell types, even differentiating between parent-daughter cells during embryogenesis. These data are a rich resource for the community that should fuel and guide future investigations into TF function. To facilitate data accessibility and utility, all strains expressing green fluorescent protein (GFP)-tagged TFs are available at the stock centers for each organism. The chromatin immunoprecipitation sequencing data are available through the ENCODE Data Coordinating Center, GEO, and through a direct interface that provides rapid access to processed data sets and summary analyses, as well as widgets to probe the cell-type-specific TF-target relationships.
© 2024 Kudron et al.; Published by Cold Spring Harbor Laboratory Press.
Figures




Update of
-
Binding profiles for 954 Drosophila and C. elegans transcription factors reveal tissue specific regulatory relationships.bioRxiv [Preprint]. 2024 Jan 20:2024.01.18.576242. doi: 10.1101/2024.01.18.576242. bioRxiv. 2024. Update in: Genome Res. 2024 Dec 23;34(12):2319-2334. doi: 10.1101/gr.279037.124. PMID: 38293065 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
