Combinatorial leaky probiotic for anticancer immunopotentiation and tumor eradication
- PMID: 39442515
- PMCID: PMC11604515
- DOI: 10.1016/j.xcrm.2024.101793
Combinatorial leaky probiotic for anticancer immunopotentiation and tumor eradication
Abstract
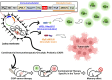
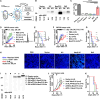
Combination therapies present a compelling therapeutic regimen against the immunosuppressive and heterogeneous microenvironment of solid tumors. However, incorporating separate therapeutic modalities in regimen designs can be encumbered by complex logistical, manufacturing, and pharmacokinetic considerations. Herein, we demonstrate a single-vector combinational anticancer therapy using an lpp gene knockout leaky probiotic for simultaneous secretion of immunotherapeutic and oncolytic effector molecules. Through fusion protein design and vector optimization, a Nissle1917 (EcN) bacteria vector is engineered to secrete Neoleukin-2/15 (Neo-2/15) cytokine-functionalized anti-PDL1 nanobody (aPDL1-Neo2/15) and anti-mesothelin-functionalized hemolysin E (HlyE-aMSLN). The multifunctional leaky probiotic enables synchronous immune activation and tumor-targeted cytolytic activity for effective tumor suppression, elevation of tumor immune cell infiltration, and establishment of anticancer immunological memory. lpp gene knockout is further shown to improve probiotic tolerability and intravenous applicability, offering a therapeutically viable approach for combination regimen development.
Keywords: anticancer immunity; bacterial therapy; chimeric protein; combinational cancer therapy; immune checkpoint inhibitors; leaky probiotic; oncolytic toxins.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures





References
-
- Moynihan K.D., Opel C.F., Szeto G.L., Tzeng A., Zhu E.F., Engreitz J.M., Williams R.T., Rakhra K., Zhang M.H., Rothschilds A.M., et al. Eradication of large established tumors in mice by combination immunotherapy that engages innate and adaptive immune responses. Nat. Med. 2016;22:1402–1410. doi: 10.1038/nm.4200. - DOI - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

