A Novel Detection and Classification Framework for Diagnosing of Cerebral Microbleeds Using Transformer and Language
- PMID: 39451369
- PMCID: PMC11504022
- DOI: 10.3390/bioengineering11100993
A Novel Detection and Classification Framework for Diagnosing of Cerebral Microbleeds Using Transformer and Language
Abstract
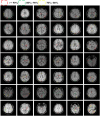
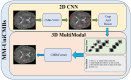
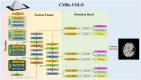
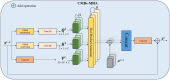
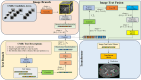
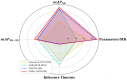
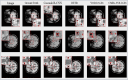
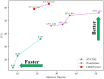
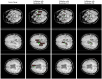
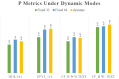
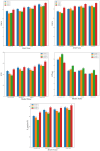
The detection of Cerebral Microbleeds (CMBs) is crucial for diagnosing cerebral small vessel disease. However, due to the small size and subtle appearance of CMBs in susceptibility-weighted imaging (SWI), manual detection is both time-consuming and labor-intensive. Meanwhile, the presence of similar-looking features in SWI images demands significant expertise from clinicians, further complicating this process. Recently, there has been a significant advancement in automated detection of CMBs using a Convolutional Neural Network (CNN) structure, aiming at enhancing diagnostic efficiency for neurologists. However, existing methods still show discrepancies when compared to the actual clinical diagnostic process. To bridge this gap, we introduce a novel multimodal detection and classification framework for CMBs' diagnosis, termed MM-UniCMBs. This framework includes a light-weight detection model and a multi-modal classification network. Specifically, we proposed a new CMBs detection network, CMBs-YOLO, designed to capture the salient features of CMBs in SWI images. Additionally, we design an innovative language-vision classification network, CMBsFormer (CF), which integrates patient textual descriptions-such as gender, age, and medical history-with image data. The MM-UniCMBs framework is designed to closely align with the diagnostic workflow of clinicians, offering greater interpretability and flexibility compared to existing methods. Extensive experimental results show that MM-UniCMBs achieves a sensitivity of 94% in CMBs' classification and can process a patient's data within 5 s.
Keywords: cerebral microbleeds; convolutional neural network; detection and classification; language–vision; multimodal.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures





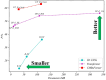

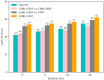
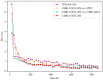
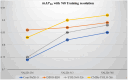

References
-
- Charidimou A., Werring D.J. Cerebral microbleeds: Detection, mechanisms and clinical challenges. Future Neurol. 2011;6:587–611. doi: 10.2217/fnl.11.42. - DOI
-
- Dora S., Kasabov N. Spiking neural networks for computational intelligence: An overview. Big Data Cogn. Comput. 2021;5:67. doi: 10.3390/bdcc5040067. - DOI
-
- Turkson R.E., Qu H., Mawuli C.B., Eghan M.J. Classification of Alzheimer’s disease using deep convolutional spiking neural network. Neural Process. Lett. 2021;53:2649–2663. doi: 10.1007/s11063-021-10514-w. - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

