Phylogenetic Analysis of Porcine Epidemic Diarrhea Virus (PEDV) during 2020-2022 and Isolation of a Variant Recombinant PEDV Strain
- PMID: 39456662
- PMCID: PMC11507624
- DOI: 10.3390/ijms252010878
Phylogenetic Analysis of Porcine Epidemic Diarrhea Virus (PEDV) during 2020-2022 and Isolation of a Variant Recombinant PEDV Strain
Abstract
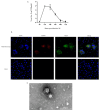
Porcine epidemic diarrhea (PED) is an acute, highly contagious, and infectious disease caused by porcine epidemic diarrhea virus (PEDV). PEDV can affect pigs of all ages, with 50~100% mortality in neonatal piglets and substantial economic losses in the swine industry. In the present study, 347 fecal and intestinal samples were collected from seven regions in China during 2020-2022. A comprehensive molecular investigation of the spike (S) gene of PEDV strains was carried out, which included phylogenetic analysis of the obtained PEDV sequences. Epidemiological surveillance data indicate that the GIIc subgroup strains are widely distributed among pigs. A PEDV strain was successfully isolated from positive small intestine samples and identified through RT-PCR detection using specific N gene primers of PEDV, indirect immunofluorescence assay (IFA), TEM analysis, genome sequencing, and full-length S gene analysis, named PEDV/SC/2022. RDP and SimPlot analysis showed that the isolate originated from the recombination of PEDV/AH2012 and PEDV/AJ1102. In conclusion, our findings contribute to the current understanding of PEDV epidemiology and provide valuable information for the control of PED outbreaks in China.
Keywords: PEDV; phylogeny; recombination; variant; virus isolation.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


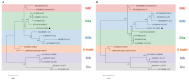


References
-
- Doyle L.P., Hutchings L.M. A transmissible gastroenteritis in pigs. J. Am. Vet. Med. Assoc. 1946;108:257–259. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

