Single Nuclei Transcriptomics Reveals Obesity-Induced Endothelial and Neurovascular Dysfunction: Implications for Cognitive Decline
- PMID: 39456952
- PMCID: PMC11508525
- DOI: 10.3390/ijms252011169
Single Nuclei Transcriptomics Reveals Obesity-Induced Endothelial and Neurovascular Dysfunction: Implications for Cognitive Decline
Abstract
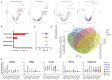
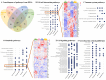
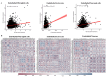
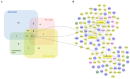
Obesity confers risk for cardiovascular disease and vascular dementia. However, genomic alterations modulated by obesity in endothelial cells in the brain and their relationship to other neurovascular unit (NVU) cells are unknown. We performed single nuclei RNA sequencing (snRNAseq) of the NVU (endothelial cells, astrocytes, microglia, and neurons) from the hippocampus of obese (ob/ob) and wild-type (WT) male mice to characterize obesity-induced transcriptomic changes in a key brain memory center and assessed blood-brain barrier permeability (BBB) by gadolinium-enhanced magnetic resonance imaging (MRI). Ob/ob mice displayed obesity, hyperinsulinemia, and impaired glucose tolerance. snRNAseq profiled 14 distinct cell types and 32 clusters within the hippocampus of ob/ob and WT mice and uncovered differentially expressed genes (DEGs) in all NVU cell types, namely, 4462 in neurons, 1386 in astrocytes, 125 in endothelial cells, and 154 in microglia. Gene ontology analysis identified important biological processes such as angiogenesis in endothelial cells and synaptic trafficking in neurons. Cellular pathway analysis included focal adhesion and insulin signaling, which were common to all NVU cell types. Correlation analysis revealed significant positive correlations between endothelial cells and other NVU cell types. Differentially expressed long non-coding RNAs (lncRNAs) were observed in cells of the NVU-affecting pathways such as TNF and mTOR. BBB permeability showed a trend toward increased signal intensity in ob/ob mice. Taken together, our study provides in-depth insight into the molecular mechanisms underlying cognitive dysfunction in obesity and may have implications for therapeutic gene targeting.
Keywords: hippocampus; neurovascular unit; obesity; single nuclei transcriptomics.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




References
-
- Ng M., Fleming T., Robinson M., Thomson B., Graetz N., Margono C., Mullany E.C., Biryukov S., Abbafati C., Abera S.F., et al. Global, regional, and national prevalence of overweight and obesity in children and adults during 1980–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet. 2014;384:766–781. doi: 10.1016/S0140-6736(14)60460-8. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

