Comprehensive Bioinformatics Analysis Reveals the Potential Role of the hsa_circ_0001081/miR-26b-5p Axis in Regulating COL15A1 and TRIB3 within Hypoxia-Induced miRNA/mRNA Networks in Glioblastoma Cells
- PMID: 39457549
- PMCID: PMC11504030
- DOI: 10.3390/biomedicines12102236
Comprehensive Bioinformatics Analysis Reveals the Potential Role of the hsa_circ_0001081/miR-26b-5p Axis in Regulating COL15A1 and TRIB3 within Hypoxia-Induced miRNA/mRNA Networks in Glioblastoma Cells
Abstract
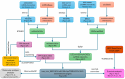
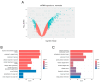
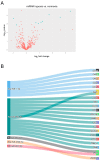
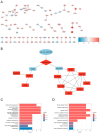
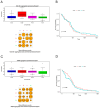
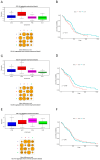
Background/Objectives: The intrinsic molecular heterogeneity of glioblastoma (GBM) is one of the main reasons for its resistance to conventional treatment. Mesenchymal GBM niches are associated with hypoxic signatures and a negative influence on patients' prognosis. To date, competing endogenous RNA (ceRNA) networks have been shown to have a broad impact on the progression of various cancers. In this study, we decided to construct hypoxia-specific microRNA/ messengerRNA (miRNA/mRNA) networks with a putative circular RNA (circRNA) regulatory component using available bioinformatics tools. Methods: For ceRNA network construction, we combined publicly available data deposited in the Gene Expression Omnibus (GEO) and interaction pairs obtained from miRTarBase and circBank; a differential expression analysis of GBM cells was performed with limma and deseq2. For the gene ontology (GO) enrichment analysis, we utilized clusterProfiler; GBM molecular subtype analysis was performed in the Glioma Bio Discovery Portal (Glioma-BioDP). Results: We observed that miR-26b-5p, generally considered a tumor suppressor, was upregulated under hypoxic conditions in U-87 MG cells. Moreover, miR-26b-5p could potentially inhibit TRIB3, a gene associated with tumor proliferation. Protein-protein interaction (PPI) network and GO enrichment analyses identified a hypoxia-specific subcluster enriched in collagen-associated terms, with six genes highly expressed in the mesenchymal glioma group. This subcluster included hsa_circ_0001081/miR-26b-5p-affected COL15A1, a gene downregulated in hypoxic U-87 MG cells yet highly expressed in the mesenchymal GBM subtype. Conclusions: The interplay between miR-26b-5p, COL15A1, and TRIB3 suggests a complex regulatory mechanism that may influence the extracellular matrix composition and the mesenchymal transformation in GBM. However, the precise impact of the hsa_circ_0001081/miR-26b-5p axis on collagen-associated processes in hypoxia-induced GBM cells remains unclear and warrants further investigation.
Keywords: GBM mesenchymal subtype; GBM molecular subtype; U-87 MG cells; bioinformatics; ceRNA; collagen; glioblastoma.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results.
Figures




References
-
- Louis D.N., Perry A., Wesseling P., Brat D.J., Cree I.A., Figarella-Branger D., Hawkins C., Ng H.K., Pfister S.M., Reifenberger G., et al. The 2021 WHO Classification of Tumors of the Central Nervous System: A summary. Neuro Oncol. 2021;23:1231–1251. doi: 10.1093/neuonc/noab106. - DOI - PMC - PubMed
-
- Torrisi F., Alberghina C., D’aprile S., Pavone A.M., Longhitano L., Giallongo S., Tibullo D., Di Rosa M., Zappalà A., Cammarata F.P., et al. The Hallmarks of Glioblastoma: Heterogeneity, Intercellular Crosstalk and Molecular Signature of Invasiveness and Progression. Biomedicines. 2022;10:806. doi: 10.3390/biomedicines10040806. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

