The Impact of Pdcd4, a Translation Inhibitor, on Drug Resistance
- PMID: 39459035
- PMCID: PMC11510623
- DOI: 10.3390/ph17101396
The Impact of Pdcd4, a Translation Inhibitor, on Drug Resistance
Abstract
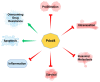
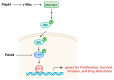
Programmed cell death 4 (Pdcd4) is a tumor suppressor, which has been demonstrated to efficiently suppress tumorigenesis. Biochemically, Pdcd4 binds with translation initiation factor 4A and represses protein translation. Beyond its role in tumor suppression, growing evidence suggests that Pdcd4 enhances the chemosensitivity of several anticancer drugs. To date, numerous translational targets of Pdcd4 have been identified. These targets govern important signal transduction pathways, and their attenuation may improve chemosensitivity or overcome drug resistance. This review will discuss the signal transduction pathways regulated by Pdcd4 and the potential mechanisms through which Pdcd4 enhances chemosensitivity or counteracts drug resistance.
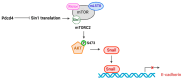
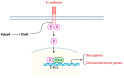
Keywords: IGF1R/IR inhibitor; JNK pathway; doxorubicin; eIF4A; fluorouracil; mTORC2–Akt pathway; paclitaxel; platinum-containing drug; β-catenin pathway.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

