Genomes of Alphanucleorhabdovirus Physostegiae Isolates from Two Different Cultivar Groups of Solanum melongena
- PMID: 39459872
- PMCID: PMC11512384
- DOI: 10.3390/v16101538
Genomes of Alphanucleorhabdovirus Physostegiae Isolates from Two Different Cultivar Groups of Solanum melongena
Abstract
Plant rhabdoviruses cause considerable economic losses and are a threat to the agriculture of Solanaceae plants. Two novel virus isolates belonging to the family Rhabdoviridae are identified by high-throughput sequencing (HTS) in Russian eggplant cultivars grown in the Volga river delta region for the first time. The phylogenetic inference of L protein (polymerase) shows that these virus isolates belong to Alphanucleorhabdovirus physostegia (Physostegia chlorotic mottle virus-PhCMoV), and their minus-sense RNA genomes have the typical gene order 3'-nucleocapsid (N)-X protein (X)-phosphoprotein (P)-Y protein (Y)-matrix protein (M)-glycoprotein (G)-polymerase (L)-5' observed in some plant-infecting alphanucleorhabdoviruses. One of the PhCMoV isolates from the eggplant cultivar Almaz is genetically very similar to the Russian PhCMoV isolate from tomato and grouped in a subclade together with four isolates from Belgium, Germany, the Netherlands, and France. However, another eggplant-infecting isolate from the Russian cultivar Boggart is the most divergent compared with the other 45 virus genomes of European PhCMoV isolates. Thus, our comparative analysis reveals that two virus isolates from Russia may either share a close evolutionary relationship with European isolates or significantly diverge from all known virus isolates. The potential to use the protein sequence comparative analysis of accessory polypeptides, along with the early developed strategy of the nucleotide sequence comparison of the RNA genomes, is shown.
Keywords: PhCMoV European isolates; Physostegia chlorotic mottle virus (PhCMoV); Russian PhCMoV isolates; high-throughput sequencing; nucleorhabdoviruses.
Conflict of interest statement
The authors declare no conflict of interests, including commercial interests. The funders had no role in the design of this study; the collection, analyses, or interpretation of data; the writing of the manuscript; or the decision to publish the results.
Figures
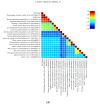





References
Publication types
MeSH terms
Substances
Associated data
- BioProject/PRJNA1144656
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

