This is a preprint.
An Extensive Atlas of Proteome and Phosphoproteome Turnover Across Mouse Tissues and Brain Regions
- PMID: 39464138
- PMCID: PMC11507808
- DOI: 10.1101/2024.10.15.618303
An Extensive Atlas of Proteome and Phosphoproteome Turnover Across Mouse Tissues and Brain Regions
Update in
-
Turnover atlas of proteome and phosphoproteome across mouse tissues and brain regions.Cell. 2025 Apr 17;188(8):2267-2287.e21. doi: 10.1016/j.cell.2025.02.021. Epub 2025 Mar 20. Cell. 2025. PMID: 40118046
Abstract
Understanding how proteins in different mammalian tissues are regulated is central to biology. Protein abundance, turnover, and post-translational modifications like phosphorylation, are key factors that determine tissue-specific proteome properties. However, these properties are challenging to study across tissues and remain poorly understood. Here, we present Turnover-PPT, a comprehensive resource mapping the abundance and lifetime of 11,000 proteins and 40,000 phosphosites across eight mouse tissues and various brain regions, using advanced proteomics and stable isotope labeling. We revealed tissue-specific short- and long-lived proteins, strong correlations between interacting protein lifetimes, and distinct impacts of phosphorylation on protein turnover. Notably, we discovered that peroxisomes are regulated by protein turnover across tissues, and that phosphorylation regulates the stability of neurodegeneration-related proteins, such as Tau and α-synuclein. Thus, Turnover-PPT provides new fundamental insights into protein stability, tissue dynamic proteotypes, and the role of protein phosphorylation, and is accessible via an interactive web-based portal at https://yslproteomics.shinyapps.io/tissuePPT.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures





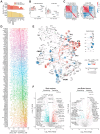

References
-
- Glass R.D., and Doyle D. (1972). On the measurement of protein turnover in animal cells. J Biol Chem 247, 5234–5242. - PubMed
-
- Pratt J.M., Petty J., Riba-Garcia I., Robertson D.H., Gaskell S.J., Oliver S.G., and Beynon R.J. (2002). Dynamics of protein turnover, a missing dimension in proteomics. Molecular & cellular proteomics : MCP 1, 579–591. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
