Conserved residues of the immunosuppressive domain of MLV are essential for regulating the fusion-critical SU-TM disulfide bond
- PMID: 39470209
- PMCID: PMC11575397
- DOI: 10.1128/jvi.00989-24
Conserved residues of the immunosuppressive domain of MLV are essential for regulating the fusion-critical SU-TM disulfide bond
Abstract
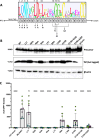
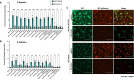
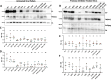
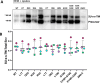
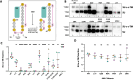
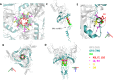
The Env protein of murine leukemia virus (MLV) is the prototype of a large clade of retroviral fusogens, collectively known as gamma-type Envs. Gamma-type Envs are found in retroviruses and endogenous retroviruses (ERVs) representing a broad range of vertebrate hosts. All gamma-type Envs contain a highly conserved stretch of 26-residues in the transmembrane subunit (TM) comprising two motifs, a putative immunosuppressive domain (ISD) and a CX6CC motif. Extraordinary conservation of the ISD and its invariant association with the CX6CC suggests a fundamental contribution to Env function. To investigate ISD function, we characterized several mutants with single amino acid substitutions at conserved positions in the MLV ISD. A majority abolished infectivity, although we did not observe a corresponding loss in intrinsic ability to mediate membrane fusion. Ratios of the surface subunit (SU) to capsid protein (CA) in virions were diminished for a majority of the ISD mutants, while TM:CA ratios were similar to wild type. Specific loss of SU reflected premature isomerization of the labile disulfide bond that links SU and TM prior to fusion. Indeed, all non-infectious mutants displayed significantly lower disulfide stability than wild-type Env. These results reveal a role for ISD positions 2, 3, 4, 7, and 10 in regulating a late step in entry after fusion peptide insertion but prior to creation of the fusion pore. This implies that the ISD is part of a larger domain, comprising the ISD and CX6CC motifs, that is critical for the formation and regulation of the metastable, intersubunit disulfide bond.IMPORTANCEThe gamma-type Env is a prevalent viral fusogen, found within retroviruses and endogenous retroviruses across vertebrate species and in filoviruses such as Ebolavirus. The fusion mechanism of gamma-type Envs is unique from other Class I fusogens such as those of influenza A virus and HIV-1. Gamma-type Envs contain a hallmark feature known as the immunosuppressive domain (ISD) that has been the subject of some controversy in the literature surrounding its putative immunosuppressive effects. Despite the distinctive conservation of the ISD, little has been done to investigate the role of this region for the function of this widespread fusogen. Our work demonstrates the importance of the ISD for the function of gamma-type Envs in infection, particularly in regulating the intermediate steps of membrane fusion. Understanding the fusion mechanism of gamma-type Envs has broad implications for understanding the entry of extant viruses and aspects of host biology connected to co-opted endogenous gamma-type Envs.
Keywords: envelope glycoprotein; fusion; gamma-type Env; immunosuppressive domain; infectivity; murine leukemia virus.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Mutation of the Putative Immunosuppressive Domain of the Retroviral Envelope Glycoprotein Compromises Infectivity.J Virol. 2017 Oct 13;91(21):e01152-17. doi: 10.1128/JVI.01152-17. Print 2017 Nov 1. J Virol. 2017. PMID: 28814524 Free PMC article.
-
Unique Structure and Distinctive Properties of the Ancient and Ubiquitous Gamma-Type Envelope Glycoprotein.Viruses. 2023 Jan 18;15(2):274. doi: 10.3390/v15020274. Viruses. 2023. PMID: 36851488 Free PMC article. Review.
-
Dysfunction of bovine endogenous retrovirus K2 envelope glycoprotein is related to unsuccessful intracellular trafficking.J Virol. 2014 Jun;88(12):6896-905. doi: 10.1128/JVI.00288-14. Epub 2014 Apr 2. J Virol. 2014. PMID: 24696495 Free PMC article.
-
Kinetic analyses of the surface-transmembrane disulfide bond isomerization-controlled fusion activation pathway in Moloney murine leukemia virus.J Virol. 2005 Nov;79(22):13856-64. doi: 10.1128/JVI.79.22.13856-13864.2005. J Virol. 2005. PMID: 16254321 Free PMC article.
-
The cytoplasmic tail of retroviral envelope glycoproteins.Prog Mol Biol Transl Sci. 2015;129:253-84. doi: 10.1016/bs.pmbts.2014.10.009. Epub 2014 Dec 1. Prog Mol Biol Transl Sci. 2015. PMID: 25595807 Free PMC article. Review.
Cited by
-
The prefusion structure of the HERV-K (HML-2) Env spike complex.Proc Natl Acad Sci U S A. 2025 Jul 15;122(28):e2505505122. doi: 10.1073/pnas.2505505122. Epub 2025 Jul 9. Proc Natl Acad Sci U S A. 2025. PMID: 40632556
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

