Quantitative proteomics and multi-omics analysis identifies potential biomarkers and the underlying pathological molecular networks in Chinese patients with multiple sclerosis
- PMID: 39478468
- PMCID: PMC11526627
- DOI: 10.1186/s12883-024-03926-3
Quantitative proteomics and multi-omics analysis identifies potential biomarkers and the underlying pathological molecular networks in Chinese patients with multiple sclerosis
Abstract
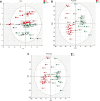
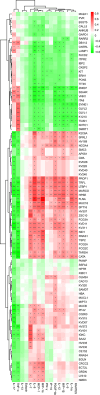
Multiple sclerosis (MS) is an autoimmune disorder caused by chronic inflammatory reactions in the central nervous system. Currently, little is known about the changes of plasma proteomic profiles in Chinese patients with MS (CpwMS) and its relationship with the altered profiles of multi-omics such as metabolomics and gut microbiome, as well as potential molecular networks that underlie the etiology of MS. To uncover the characteristics of proteomics landscape and potential multi-omics interaction networks in CpwMS, Plasma samples were collected from 22 CpwMS and 22 healthy controls (HCs) and analyzed using a Tandem Mass Tag (TMT)-based quantitative proteomics approach. Our results showed that the plasma proteomics pattern was significantly different in CpwMS compared to HCs. A total of 90 differentially expressed proteins (DEPs), such as LAMP1 and FCG2A, were identified in CpwMS plasma comparing to HCs. Furthermore, we also observed extensive and significant correlations between the altered proteomic profiles and the changes of metabolome, gut microbiome, as well as altered immunoinflammatory responses in MS-affected patients. For instance, the level of LAMP1 and ERN1 were significantly and positively correlated with the concentrations of metabolite L-glutamic acid and pro-inflammatory factor IL-17 (Padj < 0.05). However, they were negatively correlated with the amounts of other metabolites such as L-tyrosine and sphingosine 1-phosphate, as well as the concentrations of IL-8 and MIP-1α. This study outlined the underlying multi-omics integrated mechanisms that might regulate peripheral immunoinflammatory responses and MS progression. These findings are potentially helpful for developing new assisting diagnostic biomarker and therapeutic strategies for MS.
Keywords: Differentially expressed protein; Gut microbiome; Immunoinflammatory response; Metabolic profile; Multi-omics interaction networks; Multiple sclerosis; Potential biomarker; Proteomics.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures








References
-
- Yong H, Chartier G, Quandt J. Modulating inflammation and neuroprotection in multiple sclerosis. J Neurosci Res. 2018;96(6):927–50. - PubMed
-
- Sospedra M, Martin R. Immunology of multiple sclerosis. Annu Rev Immunol. 2005;23:683–747. - PubMed
-
- das Neves SP, et al. Altered astrocytic function in experimental neuroinflammation and multiple sclerosis. Glia. 2021;69(6):1341–68. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

