Targeting the hypothalamus for modeling age-related DNA methylation and developing OXT-GnRH combinational therapy against Alzheimer's disease-like pathologies in male mouse model
- PMID: 39482312
- PMCID: PMC11528003
- DOI: 10.1038/s41467-024-53507-8
Targeting the hypothalamus for modeling age-related DNA methylation and developing OXT-GnRH combinational therapy against Alzheimer's disease-like pathologies in male mouse model
Abstract
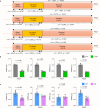
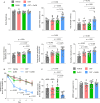
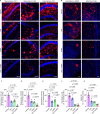
The hypothalamus plays an important role in aging, but it remains unclear regarding the underlying epigenetics and whether this hypothalamic basis can help address aging-related diseases. Here, by comparing mouse hypothalamus with two other limbic system components, we show that the hypothalamus is characterized by distinctively high-level DNA methylation during young age and by the distinct dynamics of DNA methylation and demethylation when approaching middle age. On the other hand, age-related DNA methylation in these limbic system components commonly and sensitively applies to genes in hypothalamic regulatory pathways, notably oxytocin (OXT) and gonadotropin-releasing hormone (GnRH) pathways. Middle age is associated with transcriptional declines of genes which encode OXT, GnRH and signaling components, which similarly occur in an Alzheimer's disease (AD)-like model. Therapeutically, OXT-GnRH combination is substantially more effective than individual peptides in treating AD-like disorders in male 5×FAD model. In conclusion, the hypothalamus is important for modeling age-related DNA methylation and developing hypothalamic strategies to combat AD.
© 2024. The Author(s).
Conflict of interest statement
D.C. has an invention disclosure with Albert Einstein College of Medicine (OXT-GnRH approaches for addressing AD or aging). Other authors declare no competing interests.
Figures





References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

