Integrated multi-omics revealed that dysregulated lipid metabolism played an important role in RA patients with metabolic diseases
- PMID: 39482717
- PMCID: PMC11529425
- DOI: 10.1186/s13075-024-03423-5
Integrated multi-omics revealed that dysregulated lipid metabolism played an important role in RA patients with metabolic diseases
Abstract
Objectives: Patients with rheumatoid arthritis (RA) commonly experience a high prevalence of multiple metabolic diseases (MD), leading to higher morbidity and premature mortality. Here, we aimed to investigate the pathogenesis of MD in RA patients (RA_MD) through an integrated multi-omics approach.
Methods: Fecal and blood samples were collected from a total of 181 subjects in this study for multi-omics analyses, including 16S rRNA and internally transcribed spacer (ITS) gene sequencing, metabolomics, transcriptomics, proteomics and phosphoproteomics. Spearman's correlation and protein-protein interaction networks were used to assess the multi-omics data correlations. The Least Absolute Shrinkage and Selection Operator (LASSO) machine learning algorithm were used to identify disease-specific biomarkers for RA_MD diagnosis.
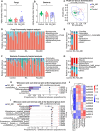
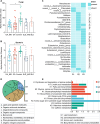
Results: Our results found that RA_MD was associated with differential abundance of gut microbiota such as Turicibacter and Neocosmospora, metabolites including decreased unsaturated fatty acid, genes related to linoleic acid metabolism and arachidonic acid metabolism, as well as downregulation of proteins and phosphoproteins involved in cholesterol metabolism. Furthermore, a multi-omics classifier differentiated RA_MD from RA with high accuracy (AUC: 0.958). Compared to gouty arthritis and systemic lupus erythematosus, dysregulation of lipid metabolism showed disease-specificity in RA_MD.
Conclusions: The integration of multi-omics data demonstrates that lipid metabolic pathways play a crucial role in RA_MD, providing the basis and direction for the prevention and early diagnosis of MD, as well as new insights to complement clinical treatment options.
Keywords: Lipid metabolism; Metabolic diseases; Multi-omics; Rheumatoid arthritis.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- McInnes IB, Schett G. The pathogenesis of rheumatoid arthritis. N Engl J Med. 2011;365(23):2205–19. - PubMed
-
- Alamanos Y, Voulgari PV, Drosos AA. Incidence and prevalence of rheumatoid arthritis, based on the 1987 American College of Rheumatology criteria: a systematic review. Semin Arthritis Rheum. 2006;36(3):182–8. - PubMed
-
- Nicolau J, et al. Rheumatoid arthritis, insulin resistance, and diabetes. Joint Bone Spine. 2017;84(4):411–6. - PubMed
-
- Del Rincon I, et al. Acceleration of atherosclerosis during the course of rheumatoid arthritis. Atherosclerosis. 2007;195(2):354–60. - PubMed
-
- Adawi M, Firas S, Blum A. Rheumatoid arthritis and atherosclerosis. Isr Med Assoc J. 2019;21(7):460–3. - PubMed
MeSH terms
Substances
Grants and funding
- 2023MS640/Administration of Traditional Chinese Medicine of Sichuan Province
- 2023MS640/Administration of Traditional Chinese Medicine of Sichuan Province
- 2021YFS0165,2022JDRC0069,22MZGC0090/Key Projects fund of the Science & Technology Department of Sichuan Province
- 21PJ085/the Health Commission of Sichuan Province
- S20001/Innovative Scientific Research Project of Medical in Sichuan Province
LinkOut - more resources
Full Text Sources
Medical

