Exploring the potential role of microbiota and metabolites in acute exacerbation of chronic obstructive pulmonary disease
- PMID: 39483760
- PMCID: PMC11526122
- DOI: 10.3389/fmicb.2024.1487393
Exploring the potential role of microbiota and metabolites in acute exacerbation of chronic obstructive pulmonary disease
Abstract
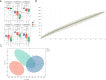
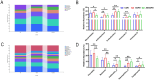
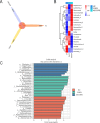
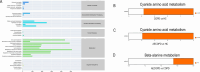
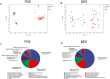
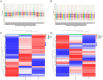
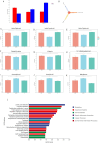
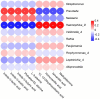
The acute exacerbation of chronic obstructive pulmonary disease seriously affects the respiratory system function and quality of life of patients. This study employed 16S rRNA sequencing and metabolomics techniques to analyze the respiratory microbiota and serum metabolites of COPD and AECOPD patients. The results showed that the microbial diversity in the respiratory tract of AECOPD patients was significantly lower than that of COPD patients, and the relative abundance of Bacteroidetes, Prevotella and Neisseria in the respiratory tract of AECOPD patients was significantly lower than that of COPD patients. However, the relative abundance of Haemophilus_D, Veillonella_A and Pseudomonas_E, in AECOPD patients was significantly higher than that of COPD patients, and the ability of respiratory microbiota in AECOPD patients to participate in alanine metabolism was significantly lower than that of COPD patients. Metabolome results further revealed that the serum alanine levels in AECOPD patients were significantly lower than those in COPD patients, and these differential metabolites were mainly involved in linoleic acid metabolism, protein digestion and absorption and regulation of lipolysis in adipocytes. In summary, the structural characteristics of respiratory microbiota in COPD and AECOPD patients are different from those in healthy populations, and their microbiota diversity decreases and microbial community structure and function will also undergo changes when acute exacerbations occur. In addition, the predicted microbial community function and metabolomics results indicate that the onset of AECOPD is mainly related to energy and amino acid metabolism disorders, especially alanine metabolism.
Keywords: AECOPD; COPD; maker; metabolite; respiratory microbiota.
Copyright © 2024 Shi, Yang, Tian, Li and Xie.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
References
LinkOut - more resources
Full Text Sources

