Lineage-specific dynamics of loss of X upregulation during inactive-X reactivation
- PMID: 39486405
- PMCID: PMC11589478
- DOI: 10.1016/j.stemcr.2024.10.001
Lineage-specific dynamics of loss of X upregulation during inactive-X reactivation
Abstract
In mammals, X chromosome dosage is balanced between sexes through the silencing of one X chromosome in females. Recent single-cell RNA sequencing analysis demonstrated that the inactivation of the X chromosome is accompanied by the upregulation of the active X chromosome (Xa) during mouse embryogenesis. Here, we have investigated if the reactivation of inactive-X (Xi) leads to the loss of Xa upregulation in different cellular or developmental contexts. We find that while Xi reactivation and loss of Xa upregulation are tightly coupled in mouse embryonic epiblast and induced pluripotent stem cells, that is not the case in germ cells. Moreover, we demonstrate that partial reactivation of Xi in mouse extra-embryonic endoderm stem cells and human B cells does not result in the loss of Xa upregulation. Finally, we have established a mathematical model for the transcriptional coordination of two X chromosomes. Together, we conclude that the reactivation of Xi is not always synchronized with the loss of Xa upregulation.
Keywords: B-cells; Epiblast; X-chromosome inactivation; X-chromosome upregulation; XEN; dosage compensation; extra-embryonic endoderm stem cells; germ cells; iPSC; pre-implantation embryos; scRNA-seq.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures
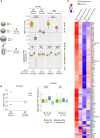




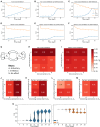
References
-
- Arava M., Majumdar S., Bammidi LS., Naik HC., Baro R., Gayen S. Gene-specific reactivation of X-linked genes upon Xist loss is linked to the chromatin states in extraembryonic endoderm and epiblast stem cells. bioRxiv. 2023 doi: 10.1101/2023.10.20.563299. Preprint at. - DOI
-
- Borensztein M., Syx L., Ancelin K., Diabangouaya P., Picard C., Liu T., Liang J.B., Vassilev I., Galupa R., Servant N., et al. Xist-dependent imprinted X inactivation and the early developmental consequences of its failure. Nat. Struct. Mol. Biol. 2017;24:226–233. doi: 10.1038/nsmb.3365. - DOI - PMC - PubMed
-
- Borensztein M., Okamoto I., Syx L., Guilbaud G., Picard C., Ancelin K., Galupa R., Diabangouaya P., Servant N., Barillot E., et al. Contribution of epigenetic landscapes and transcription factors to X-chromosome reactivation in the inner cell mass. Nat. Commun. 2017;8 doi: 10.1038/s41467-017-01415-5. - DOI - PMC - PubMed
-
- Chen G., Schell J.P., Benitez J.A., Petropoulos S., Yilmaz M., Reinius B., Alekseenko Z., Shi L., Hedlund E., Lanner F., et al. Single-cell analyses of X Chromosome inactivation dynamics and pluripotency during differentiation. Genome Res. 2016;26:1342–1354. doi: 10.1101/gr.201954.115. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

