Phospholipase A2 group IVD mediates the transacylation of glycerophospholipids and acylglycerols
- PMID: 39490928
- PMCID: PMC11621493
- DOI: 10.1016/j.jlr.2024.100685
Phospholipase A2 group IVD mediates the transacylation of glycerophospholipids and acylglycerols
Abstract
In mammalian cells, glycerolipids are mainly synthesized using acyl-CoA-dependent mechanisms. The acyl-CoA-independent transfer of fatty acids between lipids, designated as transacylation reaction, represents an additional mechanism for lipid remodeling and synthesis pathways. Here, we demonstrate that human and mouse phospholipase A2 group IVD (PLA2G4D) catalyzes transacylase reactions using both phospholipids and acylglycerols as substrates. In the presence of monoglycerol and diacylglycerol (MAG and DAG), purified PLA2G4D generates DAG and triacylglycerol, respectively. The enzyme also transfers fatty acids between phospholipids and from phospholipids to acylglycerols. Overexpression of PLA2G4D in COS7 cells enhances the incorporation of polyunsaturated fatty acids into triacylglycerol stores and induces the accumulation of lysophospholipids. In the presence of exogenously added MAG, the enzyme strongly increases cellular DAG formation, while MAG levels are decreased. PLA2G4D is not or poorly detectable in commonly used cell lines. It is expressed in keratinocytes, where it is strongly upregulated by proinflammatory cytokines. Pla2g4d-deficient mouse keratinocytes exhibit complex lipidomic changes in response to cytokine treatment, indicating that PLA2G4D is involved in the remodeling of the lipidome under inflammatory conditions. Transcriptomic analysis revealed that PLA2G4D modulates fundamental biological processes including cell proliferation, differentiation, and signaling. Together, our observations demonstrate that PLA2G4D has broad substrate specificity for fatty acid donor and acceptor lipids, allowing the acyl-CoA-independent synthesis of both phospholipids and acylglycerols. Loss-of-function studies indicate that PLA2G4D affects metabolic and signaling pathways in keratinocytes, which is associated with complex lipidomic and transcriptomic alterations.
Keywords: PLA2G4D; cPLA2δ; diacylglycerol; enzymolgy/enzyme mechanisms; glycerolipids; monoacylglycerol; phospholipases; phospholipids/metabolism; transacylase; triacylglycerol.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures
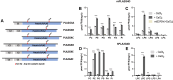








References
-
- Dodson G. Catalytic triads and their relatives. Trends Biochem. Sci. 1998;23:347–352. - PubMed
-
- Ollis D.L., Cheah E., Cygler M., Dijkstra B., Frolow F., Franken S.M., et al. The α/β hydrolase fold. Protein Eng. Des. Sel. 1992;5:197–211. - PubMed
-
- Rydel T.J., Williams J.M., Krieger E., Moshiri F., Stallings W.C., Brown S.M., et al. The crystal structure, mutagenesis, and activity studies reveal that patatin is a lipid acyl hydrolase with a ser-Asp catalytic dyad. Biochemistry. 2003;42:6696–6708. - PubMed
-
- Colaço-Gaspar M., Hofer P., Oberer M., Zechner R. PNPLA-mediated lipid hydrolysis and transacylation – at the intersection of catabolism and anabolism. Biochim. Biophys. Acta BBA Mol. Cell Biol. Lipids. 2024;1869 - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

