Oligomerization of protein arginine methyltransferase 1 and its functional impact on substrate arginine methylation
- PMID: 39491649
- PMCID: PMC11681865
- DOI: 10.1016/j.jbc.2024.107947
Oligomerization of protein arginine methyltransferase 1 and its functional impact on substrate arginine methylation
Abstract
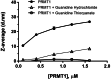
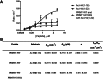
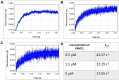
Protein arginine methyltransferases (PRMTs) are important posttranslational modifying enzymes in eukaryotic proteins and regulate diverse pathways from gene transcription, RNA splicing, and signal transduction to metabolism. Increasing evidence supports that PRMTs exhibit the capacity to form higher-order oligomeric structures, but the structural basis of PRMT oligomerization and its functional consequence are elusive. Herein, we revealed for the first time different oligomeric structural forms of the predominant arginine methyltransferase PRMT1 using cryo-EM, which included tetramer (dimer of dimers), hexamer (trimer of dimers), octamer (tetramer of dimers), decamer (pentamer of dimers), and also helical filaments. Through a host of biochemical assays, we showed that PRMT1 methyltransferase activity was substantially enhanced as a result of the high-ordered oligomerization. High-ordered oligomerization increased the catalytic turnover and the multimethylation processivity of PRMT1. Presence of a catalytically dead PRMT1 mutant also enhanced the activity of WT PRMT1, pointing out a noncatalytic role of oligomerization. Structural modeling demonstrates that oligomerization enhances substrate retention at the PRMT1 surface through electrostatic force. Our studies offered key insights into PRMT1 oligomerization and established that oligomerization constitutes a novel molecular mechanism that positively regulates the enzymatic activity of PRMTs in biology.
Keywords: PRMT1; arginine methylation; cryo-EM; enzyme activity regulation; molecular modeling; oligomerization; protein arginine methyltransferase; protein structure.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interests with the contents of this article.
Figures







References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

