Lipopeptide-mediated Cas9 RNP delivery: A promising broad therapeutic strategy for safely removing deep-intronic variants in ABCA4
- PMID: 39494150
- PMCID: PMC11531624
- DOI: 10.1016/j.omtn.2024.102345
Lipopeptide-mediated Cas9 RNP delivery: A promising broad therapeutic strategy for safely removing deep-intronic variants in ABCA4
Abstract
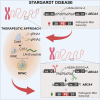
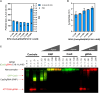
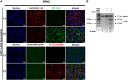
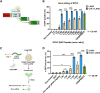
Deep-intronic (DI) variants represent approximately 10%-12% of disease-causing genetic defects in ABCA4-associated Stargardt disease (STGD1). Although many of these DI variants are amenable to antisense oligonucleotide-based splicing-modulation therapy, no treatment is currently available. These molecules are mostly variant specific, limiting their applicability to a broader patient population. In this study, we investigated the therapeutic potential of the CRISPR-Cas9 system combined with the amphipathic lipopeptide C18:1-LAH5 for intracellular delivery to correct splicing defects caused by different DI variants within the same intron. The combination of these components facilitated efficient editing of two target introns (introns 30 and 36) of ABCA4 in which several recurrent DI variants are found. The partial removal of these introns did not affect ABCA4 splicing or its expression levels when assessed in two different human cellular models: fibroblasts and induced pluripotent stem cell-derived photoreceptor precursor cells (PPCs). Furthermore, the DNA editing in STGD1 patient-derived PPCs led to a ∼50% reduction of the pseudoexon-containing transcripts resulting from the c.4539+2001G>A variant in intron 30. Overall, we provide proof-of-concept evidence of the use of C18:1-LAH5 as a delivery system for therapeutic genome editing for ABCA4-associated DI variants, offering new opportunities for clinical translation.
Keywords: ABCA4 deep-intronic variants; CRISPR-Cas9 genome editing; MT: RNA/DNA Editing; Ribonucleoprotein; Stargardt disease; intron removal; lipopeptide; peptide-mediated delivery; retina.
© 2024 The Author(s).
Conflict of interest statement
The gRNAs used for the PIR strategy for intron 30 and intron 36 as a therapeutic strategy to correct splicing defects is filed under the application no. EP24161788.5. R.W.J.C. and A.G. would like to declare that they are inventors on several patents describing the use of antisense oligonucleotides to target some of the variants in ABCA4 intron 30 and intron 36 (WO2013036105A1, WO2018109011A1, WO2020015959A1, WO2020115106A1, WO2021023863A1), which have been out-licensed to Astherna. I.V.-D. is currently partially employed by Astherna, but all work in this manuscript was performed before this appointment. R.W.J.C. is founder and Chief Scientific Officer of Astherna.
Figures



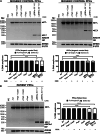

References
-
- Allikmets R., Singh N., Sun H., Shroyer N.F., Hutchinson A., Chidambaram A., Gerrard B., Baird L., Stauffer D., Peiffer A., et al. A photoreceptor cell-specific ATP-binding transporter gene (ABCR) is mutated in recessive Stargardt macular dystrophy. Nat. Genet. 1997;15:236–246. - PubMed
-
- Cremers F.P., van de Pol D.J., van Driel M., den Hollander A.I., van Haren F.J., Knoers N.V., Tijmes N., Bergen A.A., Rohrschneider K., Blankenagel A., et al. Autosomal recessive retinitis pigmentosa and cone-rod dystrophy caused by splice site mutations in the Stargardt's disease gene ABCR. Hum. Mol. Genet. 1998;7:355–362. - PubMed
-
- Lambertus S., Lindner M., Bax N.M., Mauschitz M.M., Nadal J., Schmid M., Schmitz-Valckenberg S., den Hollander A.I., Weber B.H.F., Holz F.G., et al. Progression of Late-Onset Stargardt Disease. Invest. Ophthalmol. Vis. Sci. 2016;57:5186–5191. - PubMed
-
- Huang D., Heath Jeffery R.C., Aung-Htut M.T., McLenachan S., Fletcher S., Wilton S.D., Chen F.K. Stargardt disease and progress in therapeutic strategies. Ophthalmic Genet. 2022;43:1–26. - PubMed
LinkOut - more resources
Full Text Sources

