Dilated cardiomyopathy-associated skeletal muscle actin (ACTA1) mutation R256H disrupts actin structure and function and causes cardiomyocyte hypocontractility
- PMID: 39503885
- PMCID: PMC11572969
- DOI: 10.1073/pnas.2405020121
Dilated cardiomyopathy-associated skeletal muscle actin (ACTA1) mutation R256H disrupts actin structure and function and causes cardiomyocyte hypocontractility
Abstract
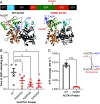
Skeletal muscle actin (ACTA1) mutations are a prevalent cause of skeletal myopathies consistent with ACTA1's high expression in skeletal muscle. Rare de novo mutations in ACTA1 associated with combined cardiac and skeletal myopathies have been reported, but ACTA1 represents only ~20% of the total actin pool in cardiomyocytes, making its role in cardiomyopathy controversial. Here we demonstrate how a mutation in an actin isoform expressed at low levels in cardiomyocytes can cause cardiomyopathy by focusing on a unique ACTA1 variant, R256H. We previously identified this variant in a family with dilated cardiomyopathy, who had reduced systolic function without clinical skeletal myopathy. Using a battery of multiscale biophysical tools, we show that R256H has potent effects on ACTA1 function at the molecular scale and in human cardiomyocytes. Importantly, we demonstrate that R256H acts in a dominant manner, where the incorporation of small amounts of mutant protein into thin filaments is sufficient to disrupt molecular contractility, and that this effect is dependent on the presence of troponin and tropomyosin. To understand the structural basis of this change in regulation, we resolved a structure of R256H filaments using cryoelectron microscopy, and we see alterations in actin's structure that have the potential to disrupt interactions with tropomyosin. Finally, we show that ACTA1R256H/+ human-induced pluripotent stem cell cardiomyocytes demonstrate reduced contractility and sarcomeric organization. Taken together, we demonstrate that R256H has multiple effects on ACTA1 function that are sufficient to cause reduced contractility and establish a likely causative relationship between ACTA1 R256H and clinical cardiomyopathy.
Keywords: actin; cardiomyopathy; contractility; muscle.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures




Update of
-
Dilated cardiomyopathy-associated skeletal muscle actin (ACTA1) mutation R256H disrupts actin structure and function and causes cardiomyocyte hypocontractility.bioRxiv [Preprint]. 2024 Mar 12:2024.03.10.583979. doi: 10.1101/2024.03.10.583979. bioRxiv. 2024. Update in: Proc Natl Acad Sci U S A. 2024 Nov 12;121(46):e2405020121. doi: 10.1073/pnas.2405020121. PMID: 38559046 Free PMC article. Updated. Preprint.
References
-
- Escobar-Lopez L., et al. , Association of genetic variants with outcomes in patients with nonischemic dilated cardiomyopathy. J. Am. Coll. Cardiol. 78, 1682–1699 (2021). - PubMed
MeSH terms
Substances
Grants and funding
- R01 GM138448/GM/NIGMS NIH HHS/United States
- T32 HL007081/HL/NHLBI NIH HHS/United States
- T32 HL007227/HL/NHLBI NIH HHS/United States
- R01HL139714/HHS | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- R01 HL139714/HL/NHLBI NIH HHS/United States
- R35 HL161185/HL/NHLBI NIH HHS/United States
- 20CVD02/Fondation Leducq (Leducq Foundation)
- R01 GM138854/GM/NIGMS NIH HHS/United States
- R01 HL151078/HL/NHLBI NIH HHS/United States
- K08 HL173569/HL/NHLBI NIH HHS/United States
- R01 HL141086/HL/NHLBI NIH HHS/United States
- 1014782/Burroughs Wellcome Fund (BWF)
- R01 HL138466/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources

