Comparative genomics of the highly halophilic Haloferacaceae
- PMID: 39506039
- PMCID: PMC11541754
- DOI: 10.1038/s41598-024-78438-8
Comparative genomics of the highly halophilic Haloferacaceae
Abstract
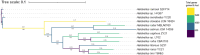
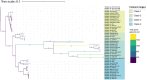
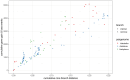
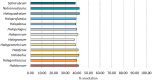
The Haloferacaceae are a family of extremely halophilic archaea with many species producing enzymes and products beneficial for industrial biotechnology. They are, however, relatively under-characterised with regards to genetics and gene products. This study aims to use existing sequence data to highlight genetic diversity, create pangenomes for three genera, and provide secondary metabolite and pathway analysis. This will establish current knowledge and identify key gaps in research. We show that the Haloferacaceae have significant genetic diversity between genera, with numerous gene gain and loss events in key genera. It also found that the model genus, Haloferax, has relatively low identified secondary metabolites compared to other genera within the family. Additionally, this study has identified potential biotechnology targets for heterologous expression in model organisms.
Keywords: Halobellus; Haloferax; Haloplanus; Archaea; Bioprospecting; Biosynthetic gene clusters; Biotechnology; Secondary metabolites.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





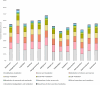
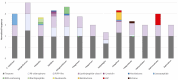
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

