ADAPT: Analysis of Microbiome Differential Abundance by Pooling Tobit Models
- PMID: 39509330
- PMCID: PMC11959182
- DOI: 10.1093/bioinformatics/btae661
ADAPT: Analysis of Microbiome Differential Abundance by Pooling Tobit Models
Abstract
Motivation: Microbiome differential abundance analysis (DAA) remains a challenging problem despite multiple methods proposed in the literature. The excessive zeros and compositionality of metagenomics data are two main challenges for DAA.
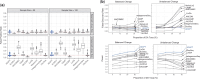
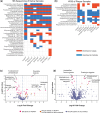
Results: We propose a novel method called "Analysis of Microbiome Differential Abundance by Pooling Tobit Models" (ADAPT) to overcome these two challenges. ADAPT interprets zero counts as left-censored observations to avoid unfounded assumptions and complex models. ADAPT also encompasses a theoretically justified way of selecting non-differentially abundant microbiome taxa as a reference to reveal differentially abundant taxa while avoiding false discoveries. We generate synthetic data using independent simulation frameworks to show that ADAPT has more consistent false discovery rate control and higher statistical power than competitors. We use ADAPT to analyze 16S rRNA sequencing of saliva samples and shotgun metagenomics sequencing of plaque samples collected from infants in the COHRA2 study. The results provide novel insights into the association between the oral microbiome and early childhood dental caries.
Availability and implementation: The R package ADAPT can be installed from Bioconductor at https://bioconductor.org/packages/release/bioc/html/ADAPT.html or from Github at https://github.com/mkbwang/ADAPT. The source codes for simulation studies and real data analysis are available at https://github.com/mkbwang/ADAPT_example.
© The Author(s) 2024. Published by Oxford University Press.
Figures

Update of
-
ADAPT: Analysis of Microbiome Differential Abundance by Pooling Tobit Models.bioRxiv [Preprint]. 2024 May 17:2024.05.14.594186. doi: 10.1101/2024.05.14.594186. bioRxiv. 2024. Update in: Bioinformatics. 2024 Nov 1;40(11):btae661. doi: 10.1093/bioinformatics/btae661. PMID: 38798558 Free PMC article. Updated. Preprint.
References
-
- Brill B, Amir A, Heller R et al. Testing for differential abundance in compositional counts data, with application to microbiome studies. Ann Appl Stat 2022;16:2648–71.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

