Organophosphorus Flame Retardants and Metabolic Disruption: An in Silico, in Vitro, and in Vivo Study Focusing on Adiponectin Receptors
- PMID: 39514743
- PMCID: PMC11548883
- DOI: 10.1289/EHP14634
Organophosphorus Flame Retardants and Metabolic Disruption: An in Silico, in Vitro, and in Vivo Study Focusing on Adiponectin Receptors
Abstract
Background: Environmental chemical exposures have been associated with metabolic outcomes, and typically, their binding to nuclear hormone receptors is considered the molecular initiating event (MIE) for a number of outcomes. However, more studies are needed to understand the influence of such exposures on cell membrane-bound adiponectin receptors (AdipoRs), which are critical metabolic regulators.
Objective: We aimed to clarify the potential interactions between AdipoRs and environmental chemicals, specifically organophosphorus flame retardants (OPFRs), and the resultant effects.
Methods: Employing in silico simulation, cell thermal shift, and noncompetitive binding assays, we screened eight OPFRs for interactions with AdipoR1 and AdipoR2. We tested two key events underlying AdipoR modulation upon OPFR exposure in a liver cell model. The Toxicological Prioritization Index (ToxPi)scoring scheme was used to rank OPFRs according to their potential to disrupt AdipoR-associated metabolism. We further examined the inhibitory effect of OPFRs on AdipoR signaling activation in mouse models.
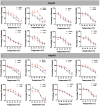
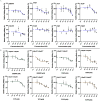
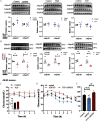
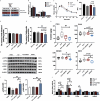
Results: Analyses identified pi-pi stacking and pi-sulfur interactions between the aryl-OPFRs 2-ethylhexyl diphenyl phosphate (EHDPP), triphenyl phosphate (TPhP), and tricresyl phosphate (TCP) and the transmembrane cavities of AdipoR1 and AdipoR2. Cell thermal shift assays showed a rightward shift in the AdipoR proteins' melting curves upon exposure to these three compounds. Although the binding sites differed from adiponectin, results suggest that aryl-OPFRs noncompetitively inhibited the binding of the endogenous peptide ligand ADP355 to the receptors. Analyses of key events underlying AdipoR modulation revealed that glucose uptake was notably lower, whereas lipid content was higher in cells exposed to aryl-OPFRs. EHDPP, TCP, and TPhP were ranked as the top three disruptors according to the ToxPi scores. A noncompetitive binding between these aryl-OPFRs and AdipoRs was also observed in wild-type (WT) mice. In db/db mice, the finding of lower blood glucose levels after ADP355 injection was diminished in the presence of a typical aryl-OPFR (TCP). WT mice exposed to TCP demonstrated lower AdipoR1 signaling, which was marked by lower phosphorylated AMP-activated protein kinase (pAMPK) and a higher expression of gluconeogenesis-related genes. Moreover, WT mice exposed to ADP355 demonstrated higher levels of pAMPK protein and peroxisome proliferator-activated receptor- messenger RNA. This was accompanied by higher glucose disposal and by lower levels of long-chain fatty acids and hepatic triglycerides; these metabolic improvements were negated upon TCP co-treatment.
Conclusions: In silico, in vitro, and in vivo assays suggest that aryl-OPFRs act as noncompetitive inhibitors of AdipoRs, preventing their activation by adiponectin, and thus function as antagonists to these receptors. Our study describes a novel MIE for chemical-induced metabolic disturbances and highlights a new pathway for environmental impact on metabolic health. https://doi.org/10.1289/EHP14634.
Figures



Similar articles
-
In vitro biolayer interferometry analysis of acetylcholinesterase as a potential target of aryl-organophosphorus flame-retardants.J Hazard Mater. 2021 May 5;409:124999. doi: 10.1016/j.jhazmat.2020.124999. Epub 2020 Dec 30. J Hazard Mater. 2021. PMID: 33454525
-
Triphenyl phosphate causes a sexually dimorphic metabolism dysfunction associated with disordered adiponectin receptors in pubertal mice.J Hazard Mater. 2020 Apr 15;388:121732. doi: 10.1016/j.jhazmat.2019.121732. Epub 2019 Nov 23. J Hazard Mater. 2020. PMID: 31796355
-
In vitro endocrine disruption potential of organophosphate flame retardants via human nuclear receptors.Toxicology. 2013 Dec 6;314(1):76-83. doi: 10.1016/j.tox.2013.09.004. Epub 2013 Sep 17. Toxicology. 2013. PMID: 24051214
-
A review of organophosphorus flame retardants (OPFRs): occurrence, bioaccumulation, toxicity, and organism exposure.Environ Sci Pollut Res Int. 2019 Aug;26(22):22126-22136. doi: 10.1007/s11356-019-05669-y. Epub 2019 Jun 26. Environ Sci Pollut Res Int. 2019. PMID: 31243659 Review.
-
Molecular mechanisms of signal transduction via adiponectin and adiponectin receptors.Biol Chem. 2010 Sep;391(9):1005-18. doi: 10.1515/BC.2010.104. Biol Chem. 2010. PMID: 20536390 Review.
Cited by
-
PKM2 regulates angiogenic activation via ANGPT2 in endothelial cells.Sci Rep. 2025 Aug 26;15(1):31448. doi: 10.1038/s41598-025-17147-2. Sci Rep. 2025. PMID: 40858916 Free PMC article.
References
-
- A O P Wiki. n.d. Welcome to the Collaborative Adverse Outcome Pathway Wiki (AOP-Wiki). https://aopwiki.org [accessed 9 July 2023].
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

