Genetic Diversity and Selection Signatures of Lvliang Black Goat Using Genome-Wide SNP Data
- PMID: 39518877
- PMCID: PMC11544794
- DOI: 10.3390/ani14213154
Genetic Diversity and Selection Signatures of Lvliang Black Goat Using Genome-Wide SNP Data
Abstract
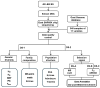
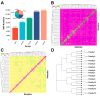
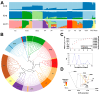
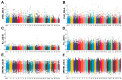
Lvliang black goat (LBG) is an excellent local breed resource in China that is known for its black fur, excellent meat quality, and strong adaptability. Studying the genetic mechanism and germplasm characteristics of LBG can provide theoretical and practical basis for the protection of the genetic resources of this breed and help implement conservation and breeding. In this study, the genetic diversity of the LBG population was evaluated using whole-genome SNP data. It was found that the LBG population had a high genetic diversity and a low degree of inbreeding. According to the clustering results of male goats and the relationship between individuals, the LBG population was divided into 13 families. Then, through population structure analysis, it was found that LBG had a close genetic relationship with the Nanjiang goat and Qinggoda goat populations, and they may have the same ancestors. The LBG population has retained some ancient genetic characteristics and is a special population that integrates local genetic characteristics and foreign gene flow. Through four selection signal analyses, we detected multiple candidate genes related to economic traits (CFL2, SCD, NLRP14, etc.) and adaptability (C4BPA, FUT8, PRNP, etc.) in the LBG population. In addition, in a comparative analysis with three commercial breeds (Saanen goat, Boer goat and Angora goat) we also found multiple genes related to physical characteristics (ERG, NRG3, EDN3, etc.). Finally, we performed functional enrichment analysis on these genes and explored their genetic mechanisms. This study provides important data support for the protection and breeding of LBG and provides a new perspective for enriching the genetic diversity of goat populations.
Keywords: Lvliang black goat; family composition; genetic diversity; population structure; selection signal.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


Similar articles
-
Whole-Genome Resequencing to Identify Selection Signatures Associated with High Fertility in Lüliang Black Goat.Animals (Basel). 2024 Dec 26;15(1):36. doi: 10.3390/ani15010036. Animals (Basel). 2024. PMID: 39794979 Free PMC article.
-
Genetic diversity and selection signatures in Hainan black goats revealed by whole-genome sequencing data.Animal. 2024 Jun;18(6):101147. doi: 10.1016/j.animal.2024.101147. Epub 2024 Mar 28. Animal. 2024. PMID: 38843669
-
Genomic insights into the conservation and population genetics of two Chinese native goat breeds.J Anim Sci. 2022 Oct 1;100(10):skac274. doi: 10.1093/jas/skac274. J Anim Sci. 2022. PMID: 35998083 Free PMC article.
-
Identifying Candidate Genes for Litter Size and Three Morphological Traits in Youzhou Dark Goats Based on Genome-Wide SNP Markers.Genes (Basel). 2023 May 29;14(6):1183. doi: 10.3390/genes14061183. Genes (Basel). 2023. PMID: 37372363 Free PMC article. Review.
-
[Progress in goat genome studies].Yi Chuan. 2019 Oct 20;41(10):928-938. doi: 10.16288/j.yczz.19-147. Yi Chuan. 2019. PMID: 31624055 Review. Chinese.
Cited by
-
Integrating SNP data to reveal the adaptive selection features of goat populations in extreme environments.BMC Genomics. 2025 Jun 2;26(1):553. doi: 10.1186/s12864-025-11743-2. BMC Genomics. 2025. PMID: 40457191 Free PMC article.
References
-
- Yayota M., Doi K. Goat Grazing for Restoring, Managing, and Conserving “Satoyama”, a Unique Socio-Ecological Production Landscape. Front. Sustain. Food Syst. 2020;4:541721. doi: 10.3389/fsufs.2020.541721. - DOI
-
- Casey N.H., Webb E.C. Managing goat production for meat quality. Small Rumin. Res. 2010;89:218–224. doi: 10.1016/j.smallrumres.2009.12.047. - DOI
-
- Wang X., Li X., Wu S., Shi K., He Y. DNA methylation and transcriptome comparative analysis for Lvliang Black goats in distinct feeding pattern reveals epigenetic basis for environment adaptation. Biotechnol. Biotechnol. Equip. 2021;35:788–795. doi: 10.1080/13102818.2021.1914164. - DOI
-
- Colli L., Milanesi M., Talenti A., Bertolini F., Chen M., Crisà A., Daly K.G., Del Corvo M., Guldbrandtsen B., Lenstra J.A., et al. Genome-wide SNP profiling of worldwide goat populations reveals strong partitioning of diversity and highlights post-domestication migration routes. Genet. Sel. Evol. 2018;50:58. doi: 10.1186/s12711-018-0422-x. - DOI - PMC - PubMed
-
- Chen N., Xia X., Hanif Q., Zhang F., Dang R., Huang B., Lyu Y., Luo X., Zhang H., Yan H., et al. Global genetic diversity, introgression, and evolutionary adaptation of indicine cattle revealed by whole genome sequencing. Nat. Commun. 2023;14:7803. doi: 10.1038/s41467-023-43626-z. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

